Int. J. Dev. Biol. 69: 195 - 202 (2025)
Dynein axonemal assembly factors (dnaaf) 5 and 9 are expressed in ciliated organs of zebrafish embryos
Open Access | Original Article | Published: 6 April 2026
Abstract
Dynein axonemal assembly factors (DNAAFs) play crucial roles in the formation and function of motile cilia, and their dysfunction often results in primary ciliary dyskinesia (PCD). We report the spatio-temporal expression patterns of dnaaf5 and dnaaf9 mRNA in zebrafish embryos, providing insight into their possible functions during development. We show that dnaaf5 and dnaaf9 mRNAs are expressed in motile ciliated tissues, such as the Kupffer’s vesicle, pronephros, floor plate, brain and olfactory placode. The dnaaf5 and dnaaf9 crispants develop ciliopathic defects during zebrafish development. These data suggest that dnaaf5 and dnaaf9 may regulate motile cilia biogenesis and function in zebrafish. Our findings suggest functional redundancy and divergence among dynein arm assembly factors in vertebrates.
Keywords
DNAAF, Primary ciliary dyskinesia, ciliogenesis, CRISPR/Cas, zebrafish
Introduction
Primary ciliary dyskinesia (PCD) is a motile ciliopathy, clinically categorised with hydrocephalus, chronic respiratory infection, infertility, situs inversus totalis or heterotaxy and hearing loss. These rare and genetically heterogeneous disorders result from mutations in genes coding for proteins mediating motile ciliogenesis and function. Structurally motile cilia possess a basic feature as primary cilia (basal body, transition zone and axonemal microtubule) and several structural augmentations to generate motion. This additional element includes a central singlet microtubule (9+2 axoneme arrangement), radial spoke, nexin link connecting the outer nine microtubule doublet with central singlet and dynein arms (inner and outer dynein arm, IDA and ODA), that function as ATP-driven motor proteins responsible for microtubule sliding and bending motions (Raidt et al., 2023). Several studies have identified that the mutations in genes associated with dynein arms assembly and trafficking cause PCD (Fassad et al., 2023, Jat et al., 2024). Axonemal dynein outer arm and inner arms are multimeric proteins that require a cytoplasmic assembly complex, which involves dynein axonemal assembly factors (DNAAF) and a transportation complex that includes the intraflagellar transport proteins.
DNAAF family proteins act as chaperone core or associated protein, facilitate assembly, stabilisation of dynein arms and sometimes their trafficking to the cilia axoneme. Like other ciliary proteins, DNAAFs are highly conserved across eukaryotes and several studies in multiple model systems, including zebrafish, have elucidated their role in motile cilia formation and function (Mitchison et al., 2012, Wan et al., 2023). These studies supported the conserved function of DNAAFs and the usefulness of zebrafish for motile cilia studies. In addition, zebrafish develop different motile ciliated organs during embryogenesis, similar to higher vertebrates, such as the brain ventricle, ependymal cell lining of the central nervous system, pronephros, Kupffer's vesicle, otic vesicle and olfactory pit. The consensus nomenclature of DNAAF lists 19 proteins under the DNAAF category (Braschi et al., 2022). Zebrafish has all orthologs except DNAAF8.
This study aims to characterise the spatiotemporal expression pattern and functional roles of dnaaf5 and dnaaf9 during early embryogenesis using zebrafish as a model organism. Zebrafish dnaaf5 and dnaaf9 show overlapping and distinct expression patterns during zebrafish embryogenesis. The expression of dnaaf5 and dnaaf9 in the motile ciliated organs during zebrafish embryogenesis is consistent with their role as dynein assembly factors.
Results
Zebrafish Dnaaf5 and Dnaaf9 show strong protein sequence conservation with human homologues
To assess the evolutionary relationship between zebrafish and human DNAAF5 and DNAAF9 homologues, their amino acid sequences were compared. DNAAF5 has a key functional domain called HEAT repeat (PF24573), which is conserved in zebrafish Dnaaf5 (Fig. 1A). Human DNAAF9 has a shulin_C20orf194-like domain (cd22936), which is also present in zebrafish Dnaaf9 (Fig. 1B).
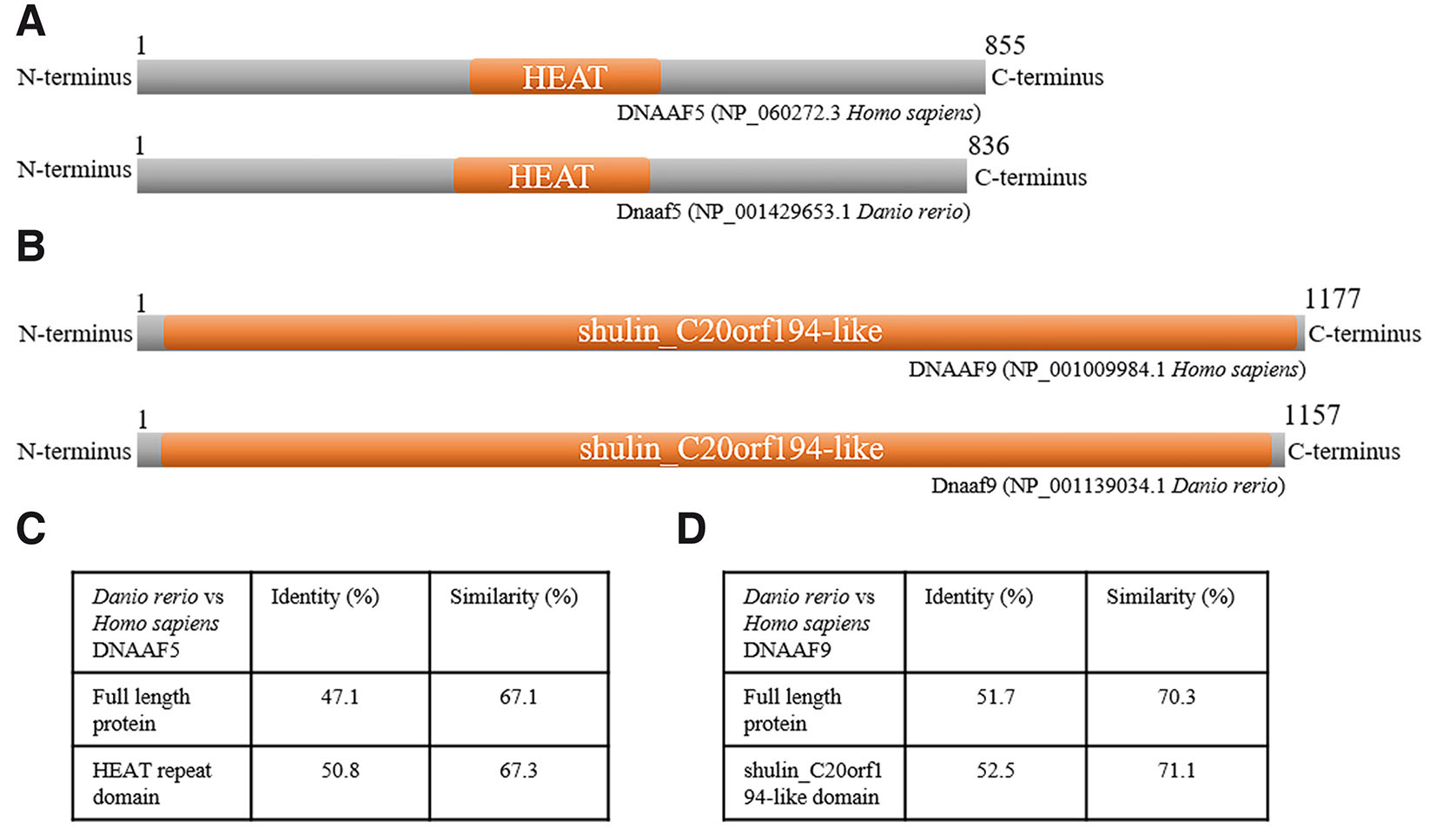
Fig. 1. Human and zebrafish DNAAF5 and DNAAF9 show conserved amino acid sequence and domain architecture.
(A,B) Schematic representation of the protein marking the HEAT domain in human and zebrafish DNAAF5 and the shulin_C20orf194-like domain in human and zebrafish DNAAF9. The length of the schematic representation is proportional to the number of amino acids in the proteins. (C,D) Tables summarising the percentage sequence identity and similarity between human and zebrafish DNAAF5 and DNAAF9 full-length protein and active domain, respectively.
We performed pairwise protein sequence alignments using EMBOSS Needle to measure the conserveness of the proteins and found that zebrafish Dnaaf5 exhibited a high degree of conservation with human DNAAF5, with 47.1% identity and 67.1% amino acid similarity across the full-length protein (Fig. 1C). The human HEAT domain shares 50.8% identity and 67.3% similarity with the zebrafish HEAT domain (Fig. 1C). The zebrafish ortholog Dnaaf9 also showed substantial conservation, sharing 51.7% identity and 70.3% similarity with human DNAAF9, while the functional domains shulin_C20orf194-like are highly conserved, showing 52.5% identity and 71.1% similarity with the respective domain of human protein (Fig. 1D). These findings indicate that both zebrafish proteins retain essential structural features of human DNAAF5 and DNAAF9, supporting their functional relevance for modelling PCD-associated gene disruption.
dnaaf5 and dnaaf9 are expressed in motile cilia-bearing organs during zebrafish development
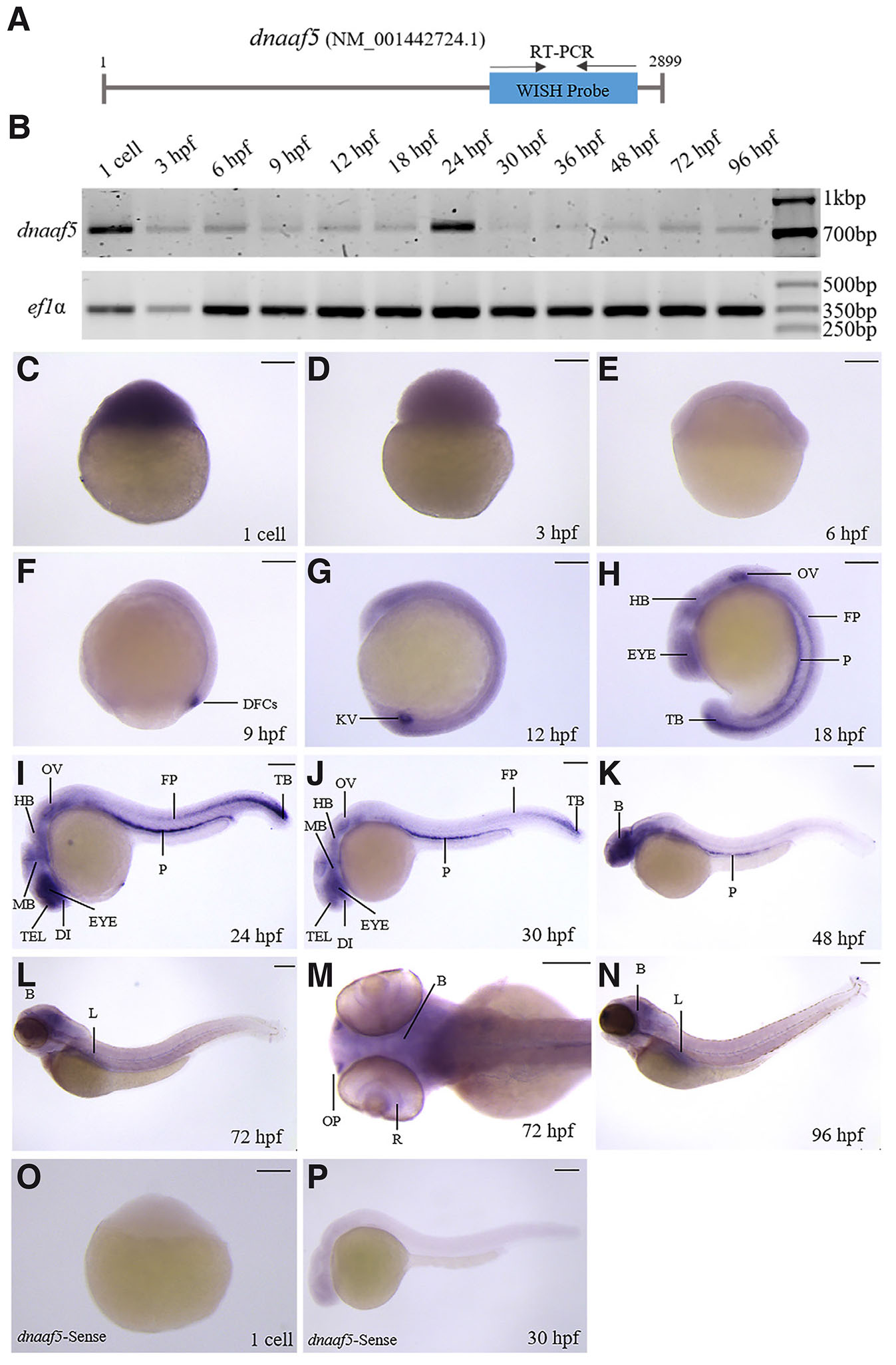
To investigate the developmental expression dynamics of dnaaf5 and dnaaf9, we carried out semi-quantitative RT-PCR and whole-mount in situ hybridisation (WISH) across multiple developmental stages of zebrafish. A 756 bp fragment of dnaaf5 was used as a WISH probe, and the primers used to amplify this fragment were used in semiquantitative RT-PCR (Fig. 2A). RT-PCR results revealed that dnaaf5 is expressed throughout zebrafish embryogenesis (Fig. 2B). WISH showed that dnaaf5 is maternally deposited and ubiquitously expressed during early embryogenesis (1-cell to 6 hpf) (Fig. 2 C-E). Thereafter, organ-specific expression of dnaaf5 can be seen (Fig. 2 F-N). It is detected in the dorsal forerunner cells (DFCs) at 9 hpf and, in a ciliated, vesicle-like organ, Kupffer's vesicle (KV) at 12 hpf (Fig. 2 F-G). Its expression can be seen in the eye, hind brain (HB) rhombomere, otic vesicle (OV), floorplate (FP), pronephros (P) and tail bud (TB) in 18 hpf embryos (Fig. 2 H-J). By 24 hpf, expression was evident in the telencephalon (TEL), diencephalon (DI), and midbrain (MB) (Fig. 2 I-J). At 48, 72 and 96 hpf, the dnaaf5 is highly expressed in the brain compared to other organs (Fig. 2 K-N). It is also found to be expressed in the liver (L) (Fig. 2 L,N). In the dorsal view of 72 hpf larvae, dnaaf5 expression can be seen in the whole brain (B), olfactory pits (OP), and retina of the eye (R) (Fig. 2M). Sense riboprobe was used as a negative control for WISH (Fig. 2 O-P).
Fig. 2. Spatiotemporal expression of dnaaf5 during zebrafish embryogenesis.
(A) Schematic representation of dnaaf5 mRNA showing the position of primers used for whole-mount in situ hybridisation (WISH) probe synthesis and RT-PCR analysis. (B) Temporal expression of dnaaf5 mRNA during zebrafish development. Zebrafish ef1α is used as a loading control. (C-N) Whole-mount in situ hybridisation showing dnaaf5 transcript localisation from the 1-cell to 96 hours post-fertilisation (hpf). (C–E) During early development (1-cell to 6 hpf), dnaaf5 is ubiquitously expressed, with strong maternal deposition of the transcript and early zygotic expression. (F-N) Later, it is expressed in specific organs. (F,G) dnaaf5 is expressed in dorsal forerunner cells (DFC) and Kupffer’s vesicle (KV) at 9 hpf and 12 hpf respectively. (H-J) 18 hpf onwards it is expressed in eye, hindbrain (HB), pronephros (P) and floorplate (FP), tail bud (TB) and otic vesicle (OV). (I-N) dnaaf5 is expressed in the telencephalon (TEL), diencephalon (DI) and midbrain (MB) after 24 hpf. (K-N) Expression is more intense in the brain than in other organs by 48 hpf. (L, N) After 72 hpf, it is expressed in the liver (L). (M) Dorsal view of 72 hpf embryo head, clarifying dnaaf5 expression throughout the brain (B), retina (R) of the eye and olfactory pit (OP). (O,P) WISH with the sense dnaaf5 probe in 1 cell and 30 hpf zebrafish embryos shows no detectable signal. Scale bars, 150 µm.
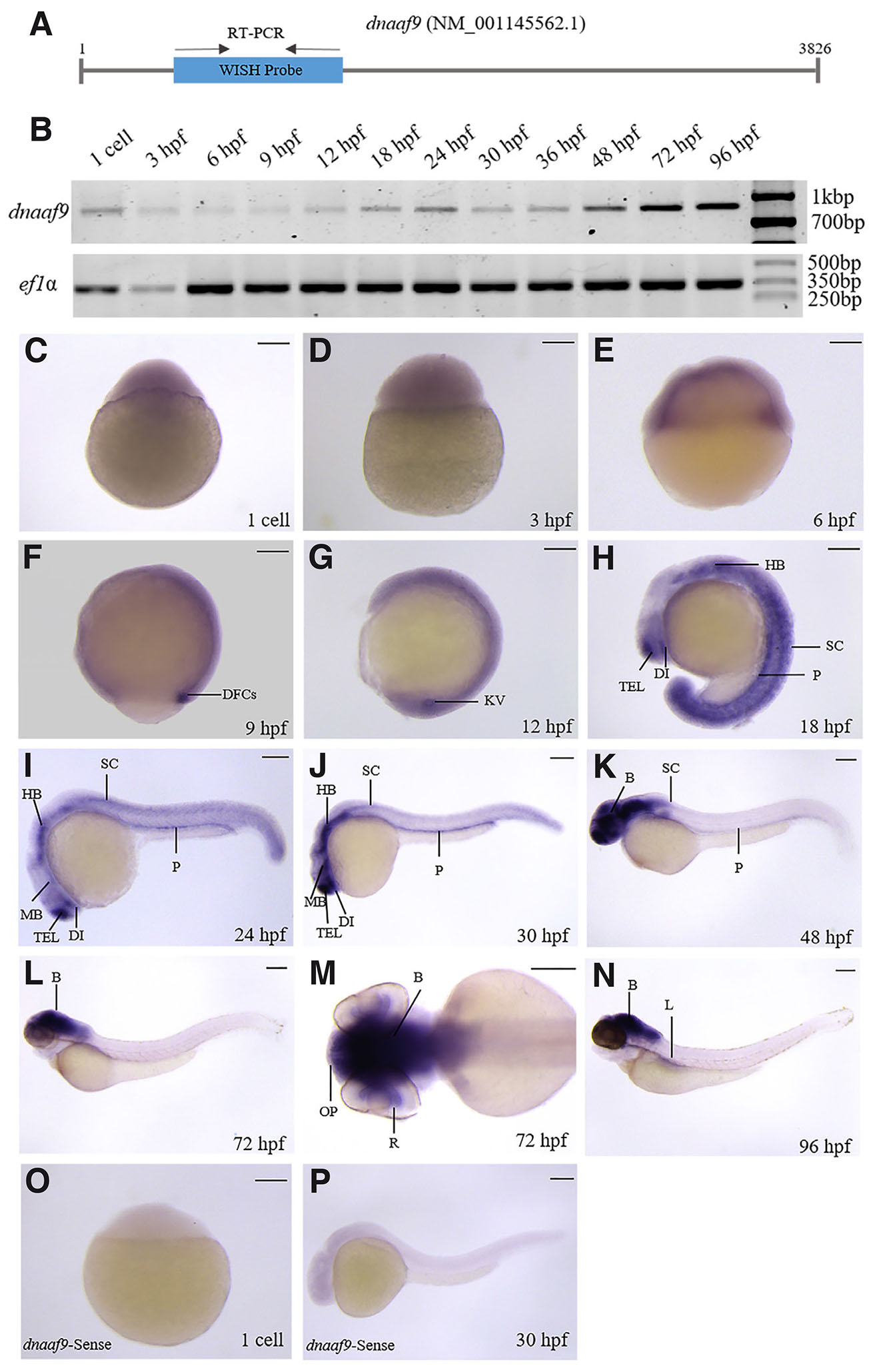
To examine the expression of dnaaf9, WISH and RT-PCR were performed using primers that span 868 bp of the transcript (Fig. 3A). Temporal expression of dnaaf9, examined by RT-PCR, shows low expression levels during developmental stages up to 48 hpf, followed by an increase thereafter (Fig. 3B). WISH revealed that the maternal deposit and early zygotic expression of dnaaf9 is minimal (Fig. 3 C-D), compared to the early gastrulation stage (Fig. 3E). Its first organ-specific expression appears at dorsal forerunner cells (DFCs), then at KV (Fig. 3 F-G). 18 hpf onwards, dnaaf9 is detected in the telencephalon (TEL), diencephalon (DI), hindbrain (HB), pronephros (P) and spinal cord (SC) (Fig. 3 H-J). After 24 hpf, expression extends to the midbrain (MB) (Fig. 3 I-N). At 48, 72 and 96 hpf, dnaaf9 is strongly expressed in the brain (B) (Fig. 3 K-N). Expression of dnaaf9 is also seen in the liver (Fig. 3N). Dorsal view of 72 hpf larvae marked dnaaf9 expression in whole brain (B), olfactory pits (OP), and retina of eye (R) (Fig. 3M). Sense riboprobe was used as a negative control for WISH (Fig. 3 O-P). These results mark the dynamic spatiotemporal expression of dnaaf5 and dnaaf9 in different ciliated organs, during zebrafish development, suggesting roles in early embryogenesis and organ-specific functions.
Fig. 3. dnaaf9 exhibits a broad expression domain across the early developmental stages of zebrafish.
(A) Schematic representation of dnaaf9 mRNA showing the position of primers used for WISH probe synthesis and RT-PCR analysis. (B) Temporal expression analysis of dnaaf9 mRNA during zebrafish development, using ef1α as a loading control. (C–N) Whole-mount in situ hybridisation marking dnaaf9 transcript localisation from the 1-cell to 96 hpf. (C–E) dnaaf9 is ubiquitously expressed during early development (1-cell to 6 hpf), with low maternal deposition and early zygotic expression. (F,G) Expression becomes restricted to dorsal forerunner cells (DFC) and Kupffer’s vesicle (KV) at 9 and 12 hpf, respectively. (H–J) From 18 hpf onward, dnaaf9 is detected in the telencephalon (TEL), diencephalon (DI), hindbrain (HB), pronephros (P) and spinal cord (SC). (I–N) At 30 hpf and later, expression extends to the midbrain (MB). (K–N) After 48 hpf, expression intensifies in the brain relative to other tissues. (N) By 96 hpf, expression is also detected in the liver (L). (M) Dorsal view of a 72 hpf embryo head showing dnaaf9 expression in the brain (B), retina (R), and olfactory pit (OP). (O,P) WISH using the dnaaf9 sense probe in 1 cell and 30 hpf zebrafish embryos shows no detectable signal. Scale bars, 150 µm.
dnaaf5 crispants exhibit a primary ciliary dyskinesia (PCD)-like phenotype
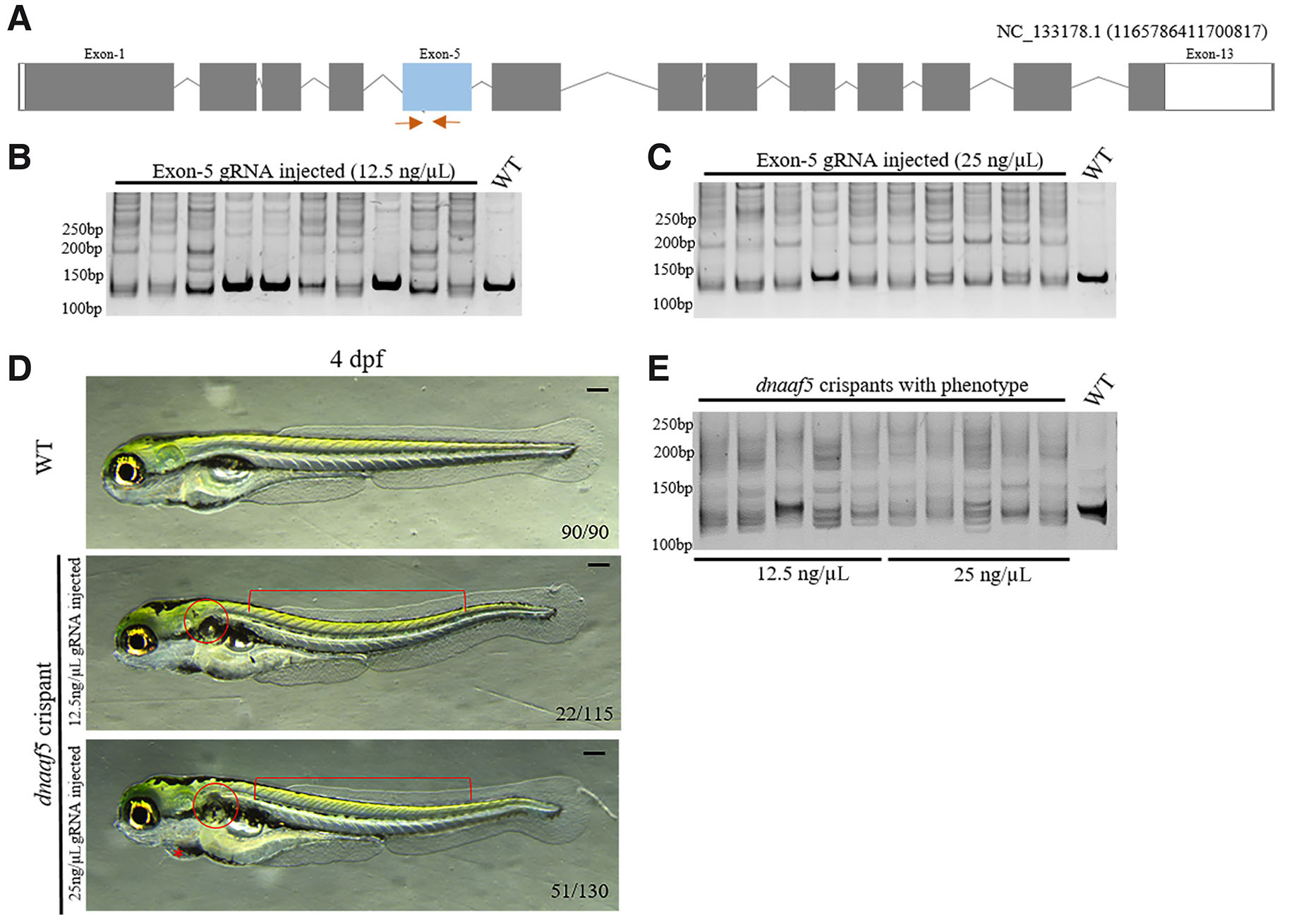
The zebrafish Dnaaf5 contains the conserved HEAT domain, spanning from 316th to 514th amino acid residue, corresponding to the coding sequences of exon 4 to exon 8 of the dnaaf5 gene. To elucidate the role of dnaaf5 in early development, we generated dnaaf5 crispants using CRISPR/Cas9 with a guide RNA (gRNA) targeting the exon 5 of dnaaf5 (Fig. 4A). Microinjection of this gRNA (12.5 pg or 25 pg per embryo) together with Cas9 mRNA into 1-cell embryos resulted in 80-90% embryos with genetic lesions (Fig. 4 B-C). The dnaaf5 crispants exhibited body curvature, glomerular cyst and pericardial edema, at different stages. The body axis curvature was observable at 2-day post fertilisation (dpf), while pericardial edema and glomerular cyst were prominent at 3 dpf. The penetrance of the phenotypes increased in a gRNA dose-dependent manner. At 12.5 pg/embryo gRNA injection, 22 out of 115 (19%) crispants showed combined glomerular cyst and body axis curvature phenotype, whereas 51 out of 130 (39%) in 25 pg/embryo gRNA injected dnaaf5 crispants exhibited the same phenotypic combination (Fig. 4D). To confirm that the phenotypes result from a mutation in the dnaaf5 gene, crispants with the phenotype were genotyped. Genotyping result confirmed the presence of mutations in the crispant showing developmental defects (Fig. 4E). Together, these results demonstrate that loss of dnaaf5 might lead to PCD-like phenotypes in zebrafish.
Fig. 4. dnaaf5 crispants exhibit primary ciliary dyskinesia (PCD)-like phenotypes.
(A) Schematic representation of the dnaaf5 gene structure, showing exons (boxes), introns (lines) and the region targeted by gRNA (blue box) along with the position of the primers used for genotyping (Orange arrow). (B,C) Genotyping results demonstrating efficient mutagenesis in 12.5 and 25 ng/μL gRNA-injected crispants. (D) Representative morphological images of 4-day post fertilisation (dpf) dnaaf5 crispants exhibiting PCD-like phenotypes, including body curvature (square grouping brace), pericardial edema (asterisk), and glomerular cyst (circle) compared to WT. (E) Genotyping result showing efficient mutation in dnaaf5 crispants showing phenotype combination. Scale bars, 150 µm.
dnaaf9 crispants exhibit scoliosis-like phenotype
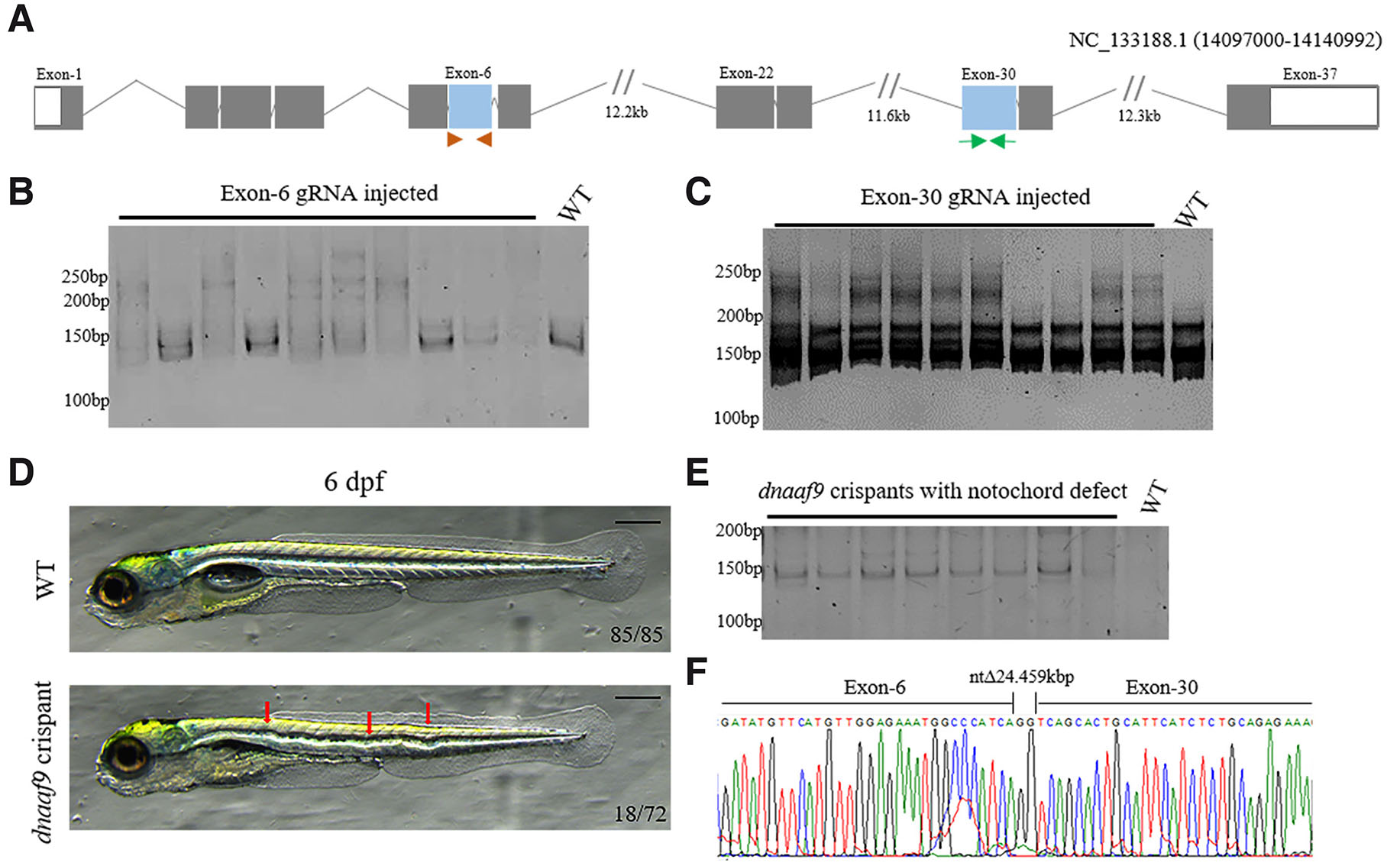
The zebrafish dnaaf9 comprises 37 exons. Exons 6 and 30 were targeted to create dnaaf9 crispant with maximal disruption of the shulin_C20orf194-like domain (Fig. 5A). Both gRNAs exhibited above 70% efficiency in inducing mutations when injected at a 25 ng/μL concentration (Fig. 5 B-C). To generate dnaaf9 crispant for phenotypic analysis, both gRNAs were co-injected into 1-cell embryos along with Cas9 mRNA. The crispants did not show morphological defects up to 3 dpf. The notochord deformities mimicking a scoliosis-like phenotype were seen in 18 out of 72 (20%) crispants by 4 to 6 dpf (Fig. 5D). To confirm that the observed phenotype resulted from the mutation of the dnaaf9 gene, crispants exhibiting the phenotype were genotyped. The PCR was performed using a forward primer from exon 6 and a reverse primer from exon 30. The expected amplicon size for wild-type DNA using these primers was 24.7 kb. In contrast, these primers will amplify a 156 bp fragment in the crispants where double-strand breaks are induced in exon 6 and exon 30, thus removing the DNA sequence between these two exons and joining them together. The PCR conditions were designed to selectively amplify the shorter fragment, while the DNA above 500 bp were not amplified. The anticipated amplicon was detected in crispants showing phenotype, while wild-type DNA failed to amplify, confirming the large deletion (Fig. 5E). To confirm the mutation, the amplified products were cloned and subjected to Sanger sequencing. Sanger sequencing confirmed deletion of a 24 kb genomic region with a small number of nucleotide insertions at the junction (Fig. 5F). This deletion corresponds to 2166 bp of the protein coding sequence, encoding 722 amino acids (residue 185 to 907), which encompasses the majority of the shulin_C20orf194-like domain, thereby confirming a functional mutation. Collectively, these results demonstrate that dnaaf9 crispants exhibit scoliosis-like phenotype in zebrafish.
Discussion
Here, we report the spatio-temporal expression pattern of dnaaf5 and dnaaf9 during zebrafish development (Figs. 2 and 3). The expression patterns of dnaaf5 and dnaaf9 in motile ciliated organs, including the KV, brain, floor plate, spinal cord, olfactory pit, and pronephros, in zebrafish embryos are consistent with the known functions of the dynein assembly factors in motile cilia. Many dnaafs exhibit a broad expression domain, while some are restricted to motile ciliated organs. Several dynein assembly factors showing maternal expression are reported (Gao et al., 2010; Panizzi et al., 2012; Falkenberg et al., 2021). wdr69 (dnaaf18) is a cilia-associated gene, expressed only in motile ciliated organs like Kupffer's vesicle, floor plate, pronephros and otic vesicle, and is functionally involved in ODA assembly. wdr69 deficient zebrafish embryos developed pronephric cyst, body curvature, pericardial edema, otolith defect, and laterality defect in different organs (Gao et al., 2010). Whereas dnaaf19 (ccdc103), along with ciliary expression, is expressed in hemato-vascular progenitors, and found to be functionally involved in myeloid cell proliferation, which is independent of cilia. Classical dynein assembly function is also exhibited by dnaaf19 (Panizzi et al., 2012; Falkenberg et al., 2021). In zebrafish, four PIH family protein genes pih1d1 (dnaaf14), pih1d2 (dnaaf15), ktu (dnaaf2), and twister (dnaaf6) are expressed in Kupffer's vesicle, floor plate, pronephros and otic vesicle, along with some parts of brains. Their functional characterisation demonstrated their importance in sperm motility and Kupffer's vesicle cilia motility, and also uncovered their role in dynein protein assembly (Yamaguchi et al., 2018). Disruption of the dyslexia candidate gene dyx1c1 (dnaaf4), expressed in KV, olfactory placode, brain, otic vesicle and pronephros, resulted in loss of cilia dynein arm and pronephric brush border defect (Chandrasekar et al., 2013). The maternal deposition of dnaaf5 and dnaaf9 expression during gastrula stages suggests that these genes may have roles during early embryogenesis.
CRISPR/Cas9-mediated F0 crispants have become valuable for preliminary characterisation of gene functions in zebrafish (Wu et al., 2018, Kroll et al., 2021). Crispants have been found to faithfully recapitulate the phenotypes seen in stable knock-out lines (Lin et al., 2025). Based on gene architecture and protein functional domain organisation, distinct CRISPR knockout strategies were employed in this study. Zebrafish dnaaf5 has a conserved HEAT domain encoded by exons 4-8. Hence, the exon 5 of this gene was targeted to disrupt the HEAT domain, thus creating a zebrafish crispant with a potentially non-functional Dnaaf5. In contrast, dnaaf9 with 37 protein-coding exons required a large fragment deletion using two gRNA targeting exon 6 and 30 to ensure major loss to the protein (Wu et al., 2018; Kim and Zhang, 2020). We observed PCD-like phenotypes in dnaaf5 crispants including pericardial edema, glomerular cyst and body curvature (Fig. 4D). These phenotypes align with previously reported phenotypes associated with mutations in human DNAAF5 (Horani et al., 2023). The dnaaf9 crispants showed scoliosis-like phenotype during late embryogenesis, with a wavy notochord (Fig. 5D). Defects in motile cilia leading to scoliosis are reported both in zebrafish and higher vertebrates (Wang et al., 2020, Marie-Hardy et al., 2021), but their underlying mechanisms are not well understood. The functional data reported here are from the F0 crispants that do not represent a homozygous mutant line. Despite this lacuna, the preliminary data presented here suggest that the zebrafish dnaaf5 and dnaaf9 knockout lines may serve as a valuable resource for investigating how defects in dynein assembly factors contribute to PCD and scoliosis.
Fig. 5. dnaaf9 crispants exhibit scoliotic-like phenotype.
>(A) Illustration of the dnaaf9 gene structure, showing exons (boxes), introns (lines) and exons targeted by guide RNAs (blue boxes) along with the primer positions used for genotyping (Orange arrowhead-Exon 6, Green arrow-Exon 30). (B,C) Heteroduplex assay result confirming high efficiency of guide RNA–mediated mutagenesis in F0 crispants. (D) Representative morphological images of 6 dpf dnaaf9 crispants exhibiting notochord malformation (red arrows). (E) Genotyping result showing efficient mutation in dnaaf9 crispants exhibiting notochord defect. (F) Representative sequencing results showing the position of the deletion of 24.7 kb between exons 6 and 30. Scale bars, 150 µm.
Materials and Methods
Zebrafish maintenance
Zebrafish were maintained in a circulating system at 28.5ºC. Embryos were raised in E3 medium at 28.5°C and staged according to Kimmel et al., (Kimmel et al., 1995). The Zebrafish Tübingen (Tü) strain was used in all experiments. All the experiments were conducted as per the approval of the Institutional Animal Ethics Committee (ILS/IAEC-250-AH/FEB-22).
Bioinformatic analysis
Zebrafish orthologue for all human DNAAFs retrieved from the ZFIN and the NCBI database. The NCBI Conserved Domain Database and NCBI ensemble were used to identify the domains present in zebrafish Dnaaf9 (NP_001139034.1), human DNAAF5 (NP_060272.3) and DNAAF9 (NP_001009984.1). The functional domain for zebrafish Dnaaf5 (NP_001429653.1) was predicted using InterProScan software. Protein sequence alignments and protein sequence identity calculation were done using EMBOSS Needle.
Gene cloning and WISH probe synthesis
Total RNA was isolated from 24 hpf zebrafish embryos using the Direct-zol RNA Miniprep kit (Zymo Research, USA). cDNA was synthesised from isolated RNA using SuperScript IV First-Strand Synthesis System (Thermo Fisher Scientific). The dnaaf5 (NM_001442724.1) and dnaaf9 (NM_001145562.1) genes were amplified by Phusion polymerase (Thermo Fisher Scientific), using gene-specific primers (Table 1), using cDNA as template. The PCR products were cloned into the PCR-Blunt-II-TOPO vector (Thermo Fisher Scientific) and were verified by Sanger sequencing. The plasmids containing the above genes were linearised with XhoI and BamHI, for antisense and sense probe synthesis, respectively. Digoxigenin-labelled antisense and sense riboprobe were synthesised by SP6 and T7 RNA polymerase.
Table 1
List of Primers
| Gene name | Forward primer | Reverse primer | Purpose | Product length |
|---|---|---|---|---|
| dnaaf5 | CTTTCCCAGCAGAGAGAGTGGT | CATCTTCAAATCTGGGAGACAAC | WISH/RT-PCR | 756 |
| dnaaf9 | ATTGTCTGACTGAAACAGAGTACGA | CCCTTATACTGAATCAACCCAAAGC | WISH/RT-PCR | 868 |
| dnaaf5 | TAAAACCCAGTGCTGTTTT | CAACCAGTCCCCAATATC | Genotyping (dnaaf5 exon 5) | 132 |
| dnaaf9 | TTGATATTTTAATAGGAC | CTTATATTTCATGGTGAA | Genotyping (dnaaf5 exon 6) |
122 |
| dnaaf9 | GGTTTGACTATGGAGCAA | GTATGTATGTATGTAAGAAG | Genotyping (dnaaf5 exon 30) |
148 |
| ef1α | CTTCTCAGGCTGACTGTGC | CCGCTAGCATTACCCTCC | RT-PCR | 358 |
Whole mount in situ hybridisation (WISH)
Embryos were treated with PTU to stop the pigmentation. For whole mount in situ hybridisation, embryos at different developmental stages were fixed in 4% paraformaldehyde (PFA), dehydrated through a graded methanol:PBST series, and stored in 100% methanol at −20 °C until use. The fixed and dehydrated embryos were rehydrated in PBST, permeabilised with proteinase-K and re-fixed in 4% PFA. Then the embryos were incubated in hybridisation buffer for 2 hours (without the probe), then the probe was added and kept at 65°C overnight. The next day, the embryos were washed with formamide/SSC buffer at 65ºC to remove unbound probes and transferred to MABT (0.1% Tween-20 in 1X MAB) at RT. Then the embryos were transferred to blocking buffer (10% fetal bovine serum (FBS) and 2% blocking reagent (Roche 1109617600) in MABT, 2 hours at RT). Then it was incubated overnight with alkaline phosphatase (AP) conjugated anti-digoxigenin (DIG) at 4ºC. On the third day, gene transcripts were detected by developing with BM purple (Roche 11442074001). The reaction was stopped by adding PBS and fixed with 4% PFA kept at 4ºC for overnight.
RT-PCR
cDNA was prepared by taking an equal amount of total RNA extracted from different developmental stages. The above-mentioned WISH primer pairs were used to amplify 756 bp and 868 bp of dnaaf5 and dnaaf9, respectively (Table 1). Zebrafish ef1alpha (now called eukaryotic translation elongation factor 1 alpha 1, like 1 (eef1a1l1), NM_131263.1) was used as a loading control.
gRNA design and synthesis
gRNA target for dnaaf5 and dnaaf9 was designed using CRISPR RGEN tools (http://www.rgenome.net/cas-designer/). gRNAs were designed against exon 5 (5’-GATTCCTTACCACCAGCTCC-3’) of dnaaf5. For dnaaf9 crispants, gRNAs were designed to target exon 6 and exon 30, having sequence 5’-GGAGAAATGGCCCATCATTC-3’ and 5’-GATGAATGCAGTGCTGAGAT-3’ respectively. DNA templates for the designed gRNA target and the universal oligo were obtained from IDT and annealed to prepare the template for gRNA transcription. gRNA synthesised from annealed template using MEGAscript SP6 or T7 kit (Thermo Fisher Scientific) (Gagnon et al., 2014).
Microinjection
Crispants were generated by injecting 1nl of gRNA and Cas9 mRNA mixture, containing each gRNA (for one gene) at a concentration of 12.5 ng/μL or 25 ng/μL and Cas9 mRNA at a concentration of 100 ng/μL. Microinjection was performed at the single cell stage of zebrafish embryos using a Femtojet microinjector (Eppendorf).
High resolution melt curve analysis (HRM) and Heteroduplex assay (HD)
Genomic DNA was isolated from the whole embryos 48 hours post gRNA/Cas9 injection in PCR tubes by incubating in TE buffer at 95°C and digested by proteinase K at 55°C (Zhu et al., 2014). For DNA amplification, primers were designed flanking the CRISPR target site (Table 1). The reaction for HRM was prepared by adding 2X SYBR® Green JumpStartTM Taq ReadyMix, genomic DNA, forward and reverse primers (10 μM) and nuclease-free water. PCR reaction method (95°C/15 sec., 50-60°C /20sec and 72°C /20 sec) was set for 40 cycles. During melt curve reaction temperature was increased from 60-95°C with an increment of 0.2°C and fluorescence was recorded at every 0.2 sec.
In the HD assay, the amplified DNA obtained from HRM was denatured at 95°C and reannealed at RT. The products were then loaded on 12% polyacrylamide gel and electrophoresis was carried out at 120V for 90 minutes, to analyse the heteroduplex and homoduplex bands formed in the crispant compared to the WT homoduplex.
Microscopy
The live zebrafish embryos were anaesthetised in Tricaine methanesulfonate solution, and then dropped on 3% methylcellulose bed. Thereafter, the bright-field images were obtained, while fixed samples were mounted in 75% glycerol and images were taken. All the imaging was taken using a Leica M205FA stereo microscope.
Acknowledgements
This work was partially supported by the DBT - mission program on pediatric rare genetic disorders (BT/PR45460/MED/12/952/2022) grant to RKS, and intramural funds from BRIC-ILS, which is an institute of the Department of Biotechnology, Government of India. UN is a recipient of the DST-Inspire fellowship (IF180156) and KS is a UGC-SRF (221610148651).
Declarations
Conflicts of interest
The authors declare no conflict of interest.
Author contributions
RKS conceived, supervised and provided resources for the study. UN and KS carried out the experiments. UN wrote the manuscript. RKS and KS revised the manuscript. All the authors approved the final version of the manuscript.
References
Braschi B., Omran H., Witman G. B., Pazour G. J., Pfister K. K., Bruford E. A., King S. M. (2022). Consensus nomenclature for dyneins and associated assembly factors. Journal of Cell Biology 221: e202109014.
Chandrasekar G., Vesterlund L., Hultenby K., Tapia-Páez I., Kere J. (2013). The Zebrafish Orthologue of the Dyslexia Candidate Gene DYX1C1 Is Essential for Cilia Growth and Function. PLoS ONE 8: e63123.
Falkenberg L. G., Beckman S. A., Ravisankar P., Dohn T. E., Waxman J. S. (2021). Ccdc103 promotes myeloid cell proliferation and migration independent of motile cilia. Disease Models & Mechanisms 14: dmm048439.
Fassad M. R., Rumman N., Junger K., Patel M. P., Thompson J., Goggin P., Ueffing M., Beyer T., Boldt K., Lucas J. S., Mitchison H. M. (2023). Defective airway intraflagellar transport underlies a combined motile and primary ciliopathy syndrome caused by IFT74 mutations . Human Molecular Genetics 32: 3090-3104.
Gagnon J. A., Valen E., Thyme S. B., Huang P., Ahkmetova L., Pauli A., Montague T. G., Zimmerman S., Richter C., Schier A. F. (2014). Efficient Mutagenesis by Cas9 Protein-Mediated Oligonucleotide Insertion and Large-Scale Assessment of Single-Guide RNAs. PLoS ONE 9: e98186.
Gao C., Wang G., Amack J. D., Mitchell D. R. (2010). Oda16/Wdr69 is essential for axonemal dynein assembly and ciliary motility during zebrafish embryogenesis. Developmental Dynamics 239: 2190-2197.
Horani A., Gupta D. K., Xu J., Xu H., Carmen Puga-Molina L. , Santi C. M., Ramagiri S., Brennan S. K., Pan J., Koenitzer J. R., Huang T., Hyland R. M., Gunsten S. P., Tzeng S.C., Strahle J. M., Mill P., Mahjoub M. R., Dutcher S. K., Brody S. L. (2023). The effect of Dnaaf5 gene dosage on primary ciliary dyskinesia phenotypes. JCI Insight 8: e168836.
Jat K. R., Faruq M., Jindal S., Bari S., Soni A., Sharma P., Mathews S., Shamim U., Ahuja V., Uppilli B., Yadav S. C., Lodha R., Arava S. K., Kabra S. K. (2024). Genetics of 67 patients of suspected primary ciliary dyskinesia from India. Clinical Genetics 106: 650-658.
Kim B. H., Zhang G.J. (2020). Generating Stable Knockout Zebrafish Lines by Deleting Large Chromosomal Fragments Using Multiple gRNAs. G3 Genes|Genomes|Genetics 10: 1029-1037.
Kimmel C. B., Ballard W. W., Kimmel S. R., Ullmann B., Schilling T. F. (1995). Stages of embryonic development of the zebrafish. Developmental Dynamics 203: 253-310.
Kroll F., Powell G. T., Ghosh M., Gestri G., Antinucci P., Hearn T. J., Tunbak H., Lim S., Dennis H. W., Fernandez J. M., Whitmore D., Dreosti E., Wilson S. W., Hoffman E. J., Rihel J. (2021). A simple and effective F0 knockout method for rapid screening of behaviour and other complex phenotypes. eLife 10: e59683.
Lin S.J., Huang K., Petree C., Qin W., Varshney P., Varshney G. K. (2025). Optimizing gRNA selection for high-penetrance F0 CRISPR screening for interrogating disease gene function. Nucleic Acids Research 53: gkaf180.
Marie-Hardy L., Cantaut-Belarif Y., Pietton R., Slimani L., Pascal-Moussellard H. (2021). The orthopedic characterization of cfap298tm304 mutants validate zebrafish to faithfully model human AIS. Scientific Reports 11: 7392.
Mitchison H. M., Schmidts M., Loges N. T., Freshour J., Dritsoula A., Hirst R. A., O'Callaghan C., Blau H., Al Dabbagh M., Olbrich H., Beales P. L., Yagi T., Mussaffi H., Chung E. M. K., Omran H., Mitchell D. R. (2012). Mutations in axonemal dynein assembly factor DNAAF3 cause primary ciliary dyskinesia. Nature Genetics 44: 381-389.
Panizzi J. R., Becker-Heck A., Castleman V. H., Al-Mutairi D. A., Liu Y., Loges N. T., Pathak N., Austin-Tse C., Sheridan E., Schmidts M., Olbrich H., Werner C., Häffner K., Hellman N., Chodhari R., Gupta A., Kramer-Zucker A., Olale F., Burdine R. D., Schier A. F., O'Callaghan C., Chung E. M. K., Reinhardt R., Mitchison H. M., King S. M., Omran H., Drummond I. A. (2012). CCDC103 mutations cause primary ciliary dyskinesia by disrupting assembly of ciliary dynein arms. Nature Genetics 44: 714-719.
Raidt J., Loges N. T., Olbrich H., Wallmeier J., Pennekamp P., Omran H. (2023). Primary ciliary dyskinesia. La Presse Médicale 52: 104171.
Wan F., Yu L., Qu X., Xia Y., Feng K., Zhang L., Zhang N., Zhao G., Zhang C., Guo H. (2023). A novel mutation in PCD ‐associated gene DNAAF3 causes male infertility due to asthenozoospermia . Journal of Cellular and Molecular Medicine 27: 3107-3116.
Wang Y., Liu Z., Yang G., Gao Q., Xiao L., Li J., Guo C., Troutwine B. R., Gray R. S., Xie L., Zhang H. (2020). Coding Variants Coupled With Rapid Modeling in Zebrafish Implicate Dynein Genes, dnaaf1 and zmynd10, as Adolescent Idiopathic Scoliosis Candidate Genes. Frontiers in Cell and Developmental Biology 8: 582255.
Wu R. S., Lam I. I., Clay H., Duong D. N., Deo R. C., Coughlin S. R. (2018). A Rapid Method for Directed Gene Knockout for Screening in G0 Zebrafish. Developmental Cell 46: 112-125.e4.
Yamaguchi H., Oda T., Kikkawa M., Takeda H., (2018). Systematic studies of all PIH proteins in zebrafish reveal their distinct roles in axonemal dynein assembly. eLife 7: e36979.
Zhu X., Xu Y., Yu S., Lu L., Ding M., Cheng J., Song G., Gao X., Yao L., Fan D., Meng S., Zhang X., Hu S., Tian Y. (2014). An Efficient Genotyping Method for Genome-modified Animals and Human Cells Generated with CRISPR/Cas9 System. Scientific Reports 4: 6420.