Int. J. Dev. Biol. 69: 187 - 194 (2025)
A chloroquine sensitivity gradient induces tissue regeneration and maintenance phenotypes in planarians
Open Access | Original Article | Published: 6 April 2026
Abstract
Failure to properly regenerate and maintain tissues is a root cause of disease and aging. One goal of regenerative biology is to unveil mechanisms by which simpler model organisms successfully regenerate so that we can better understand how to control the cellular and molecular switches in human tissues. Planarians have become a favored model system for these studies because of their amazing capacity to fully and rapidly regenerate following amputation, but also for their response to other stressors such as starvation. Planarians undergo proportional degrowth when starved, a unique process that balances stem cell proliferation and apoptotic cell death, autophagy-mediated cell turnover, and energy sensing. We chose a pharmacological approach to study autophagy in planarians by using chloroquine, an autophagy inhibitor. We found that head regression and tissue lysis were induced by chloroquine in whole planarians and that starved planarians were more sensitive than their fed counterparts. While chloroquine treatment did not prevent tissue regeneration following injury, it caused regeneration delays and disruptions in photoreceptor replacement. Finally, we show that an anterior-posterior gradient of chloroquine sensitivity exists in the planarian with prepharyngeal tissues exhibiting significantly low sensitivity. Together, our results extend our understanding of autophagic flux in planarian tissue regeneration and maintenance and establish a model by which an autophagic gradient exists in this highly regenerative system.
Keywords
autophagy, chloroquine, gradient, neoblast, planarian, regeneration, stem cell
Introduction
The ability to maintain normal cell function and tissue morphology is critical for homeostatic body maintenance and failure to do so often leads to disease and death. While humans and other mammals are notorious for their inability to regenerate lost or damaged tissue, some organisms exhibit a strong capacity to regenerate complex structures (Poss and Tanaka, 2024). One of the most amazing examples of tissue regeneration is observed in freshwater planarians. Planarians are flatworms that can regenerate every cell type even from tiny amputated fragments (Reddien and Sánchez Alvarado, 2004). A population of pluripotent stem cells, called neoblasts, respond to injury by proliferating and migrating to the injury site, thus giving rise to a wound-adjacent accumulation of mitotically active neoblasts (Newmark and Sánchez Alvarado, 2000; Sahu et al., 2021; Wagner et al., 2011; Wenemoser and Reddien, 2010). Daughter cells of these mitoses contribute to a growing blastema, a structure observed in other regenerative model organisms (Eisenhoffer et al., 2008; Poss and Tanaka, 2024). Neoblasts receive muscle-derived positional guidance cues to differentiate into the missing cell types giving rise to a fully regenerated animal that is correctly patterned along all body axes (Adell et al., 2010; Scimone et al., 2017; Witchley et al., 2013).
A remarkable characteristic of planarian regeneration is that it can occur in tiny fragments despite the removal of a muscular pharynx and an organized intestinal system, two structures that are critical for the intake, digestion, and distribution of external nutrients. The planarian undergoes a poorly understood tissue remodeling process (morphallaxis) that reestablishes a normal body plan containing pharyngeal and digestive tissues, allowing for food intake and subsequent organismal growth (Pellettieri, 2019; Reddien and Sánchez Alvarado, 2004). Planarian tissues are constantly and rapidly undergoing turnover events that are balanced by the production of new neoblast-derived cells (Lee et al., 2024; Lindsay-Mosher et al., 2024). These tissue remodeling events work in conjunction with the proliferation of neoblasts to seamlessly blend new and pre-existing tissues. Remodeling extends to physiological tissue maintenance in whole unamputated planarians. Planarians respond to long periods of starvation by degrowing, a process partially driven by apoptotic cell death (Adell et al., 2025; Pellettieri et al., 2010). This unique response to starvation can continue for long periods of time as the planarian maintains proper body proportion and morphology (Emili et al., 2025; Takeda et al., 2009; Thommen et al., 2019). Therefore, planarians exhibit remarkable capacity to endogenously support the energy requirements of stem cell-based tissue regeneration and maintenance and establish themselves as excellent models to understand the cellular and molecular signals that govern these events.
This unique level of tissue plasticity underlies the planarian’s ability to successfully respond to a variety of external stressors such as tissue loss and starvation and suggests that planarians maintain their tissues through a strong cellular autophagy mechanism. Autophagy is an endogenous process by which cells degrade and recycle intracellular materials including proteins and organelles (Yamamoto et al., 2023). This catabolic breakdown of cellular components provides the cell with energy and recycled building blocks for future biosynthesis and allows for cell survival during periods of stress, including starvation. Autophagy is initiated when upstream stress responses trigger the nucleation and elongation of a phagophore membrane that engulfs cellular materials to form a vesicular structure called the autophagosome. The autophagosome subsequently fuses with the lysosome whereby hydrolytic enzymes degrade the delivered cargo (Ballabio and Bonifacino, 2020).
Autophagic flux remains critical for normal cellular physiology and homeostasis as studies show that knockdown of autophagy regulators leads to embryonic dysmorphias and lethality (Allen and Baehrecke, 2020), pathologies including neurodegenerative diseases and cancer (Aman et al., 2021), and increased or decreased lifespan (Hansen et al., 2018). In support of cellular plasticity, autophagy has been shown to play important roles in regulating stem cell pluripotency, self-renewal, and differentiation (Boya et al., 2018; Tang and Rando, 2014). Tissue regeneration studies involving the targeted knockdown of autophagy genes have highlighted a conserved role for autophagy in zebrafish caudal fin and extraocular muscle regeneration (Saera-Vila et al., 2016; Varga et al., 2014), intestinal stem cell proliferation in Drosophila digestive system maintenance (Nagy et al., 2018), and mammalian keratinocyte proliferation, migration, and differentiation during epidermal wound healing (Du et al., 2024; Qiang et al., 2021). Some studies have taken pharmacological approaches to modify autophagic flux. Exogenous treatment with chemicals known to inhibit autophagy impairs head regeneration in hydra (Suknovic et al., 2021), mesentery regeneration in sea anemones (Bossert and Thomsen, 2017), fin regeneration in zebrafish (Chen et al., 2024), cell proliferation and blastema formation in injured newt limbs (Peng et al., 2021), and muscle and nerve regeneration in rodents (Nichenko et al., 2016). Pharmacological approaches to induce autophagy appear to improve nerve, muscle, and liver regeneration in rodents (Balnis et al., 2025; Ding et al., 2018; Huang et al., 2016; Xiang et al., 2021) and heart regeneration in zebrafish (Xie et al., 2021). Together, these suggest that upregulation of autophagy contributes to normal tissue regeneration and maintenance in a variety of model systems, and loss of autophagic flux leads to a failure to regenerate tissues upon damage.
Several studies have implicated autophagy in planarian tissue maintenance beginning with the discovery that the ortholog for human death-associated protein-1 (Gtdap-1) is upregulated after injury or starvation and RNA interference (RNAi)-based knockdown of its function leads to delayed regeneration and tissue lysis (González-Estévez et al., 2007a, 2007b). Planarian studies focusing on other autophagy-related genes (atg) unveil similar injury- and starvation-induced expression, dysmorphic blastemal tissue, decreased regeneration speed, and head regression phenotypes (Jin et al., 2022; Kang et al., 2019; Ma et al., 2025; Ma et al., 2018). Furthermore, proteins that sense the status of cell energy have been implicated in balancing the mitotic/apoptotic ratio that controls tissue maintenance and growth after injury and starvation (González-Estévez et al., 2012; Oviedo et al., 2008; Pascual-Carreras et al., 2020; Tu et al., 2012). The planarian is an excellent model through which to study the role of autophagy because of its ability to fully regenerate tissue and to transition from growth to proportional degrowth. In our study, we take a pharmacological approach to characterizing the role of autophagy in Schmidtea mediterranea. We find that treatment with a known late-stage autophagy inhibitor, chloroquine, results in planarian head regression and tissue lysis and that starved planarians are significantly more sensitive compared to their fed counterparts. Following injury, chloroquine reduces regeneration speed and negatively impacts photoreceptor regeneration but does not completely inhibit new tissue replacement. We also find that a gradient of chloroquine sensitivity exists in the planarian as different regenerates along the anterior-posterior (A/P) axis undergo tissue lysis over time in a tail-to-prepharyngeal sequence with prepharyngeal tissues exhibiting significantly low sensitivity. Our results highlight the importance of autophagy-based turnover in homeostatic tissue maintenance in the planarian, suggest a potential underlying autophagy gradient, and open new paths to studying the effects of chloroquine in a highly regenerative model system.
Results and Discussion
Chloroquine induces Planarian head regression and lysis
Chloroquine is a lysosomotropic compound that inhibits autophagic flux by increasing lysosomal pH, thus blocking autophagosome-lysosome fusion and inhibiting hydrolytic enzyme function (Mauthe et al., 2018). We chose a chloroquine-based strategy to study autophagy and lysosomal function for several reasons. Chloroquine diphosphate water solubility allows for direct planarian treatment in culture and removes the need for a feeding- or injection-based administration of double-stranded RNA (dsRNA) for RNAi-based knockdown of autophagy-related genes. This approach presumably provides ubiquitous and sustained autophagy inhibition in all cells as RNAi approaches are tied to genes expressed only in specific cell types.
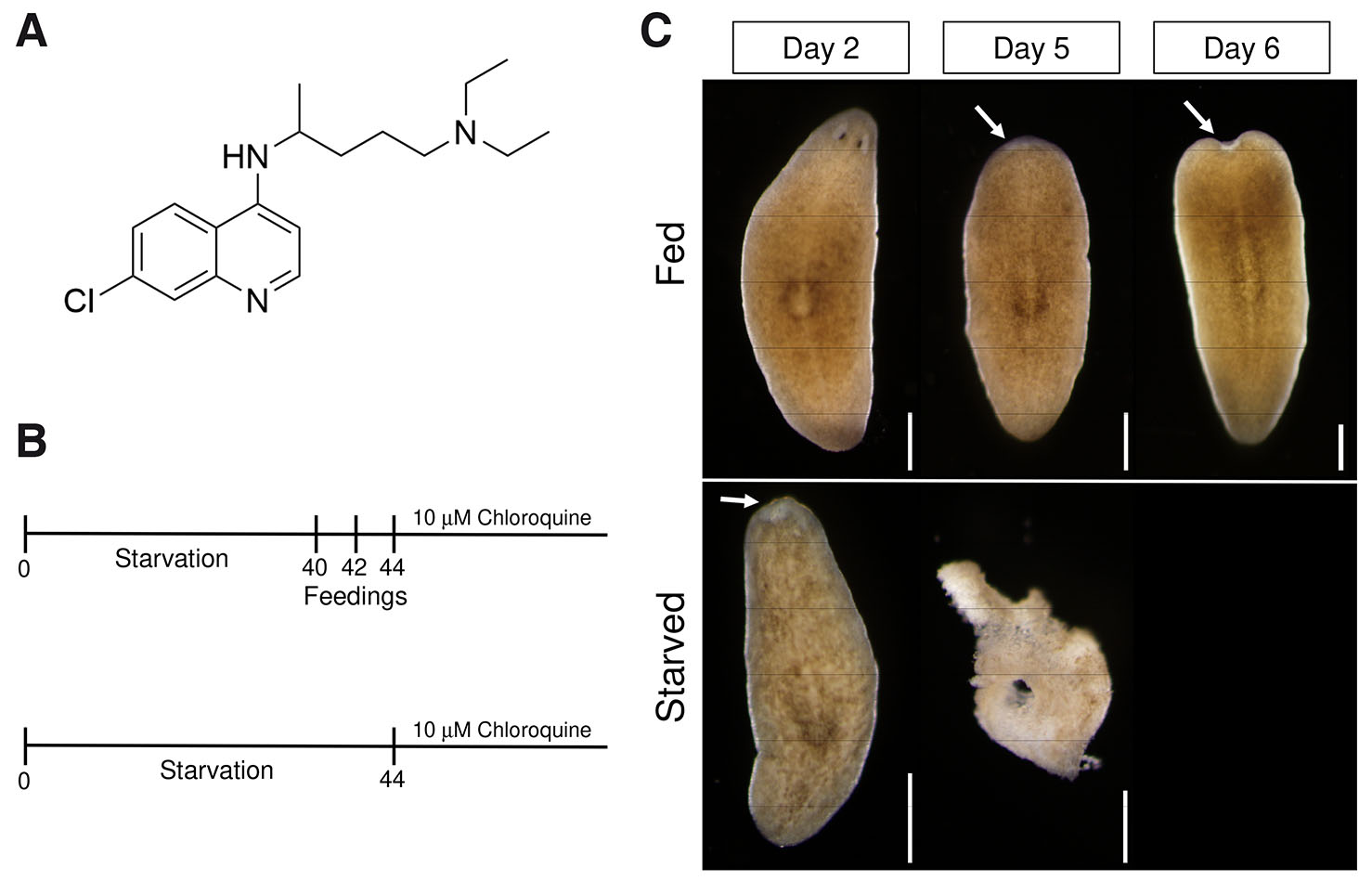
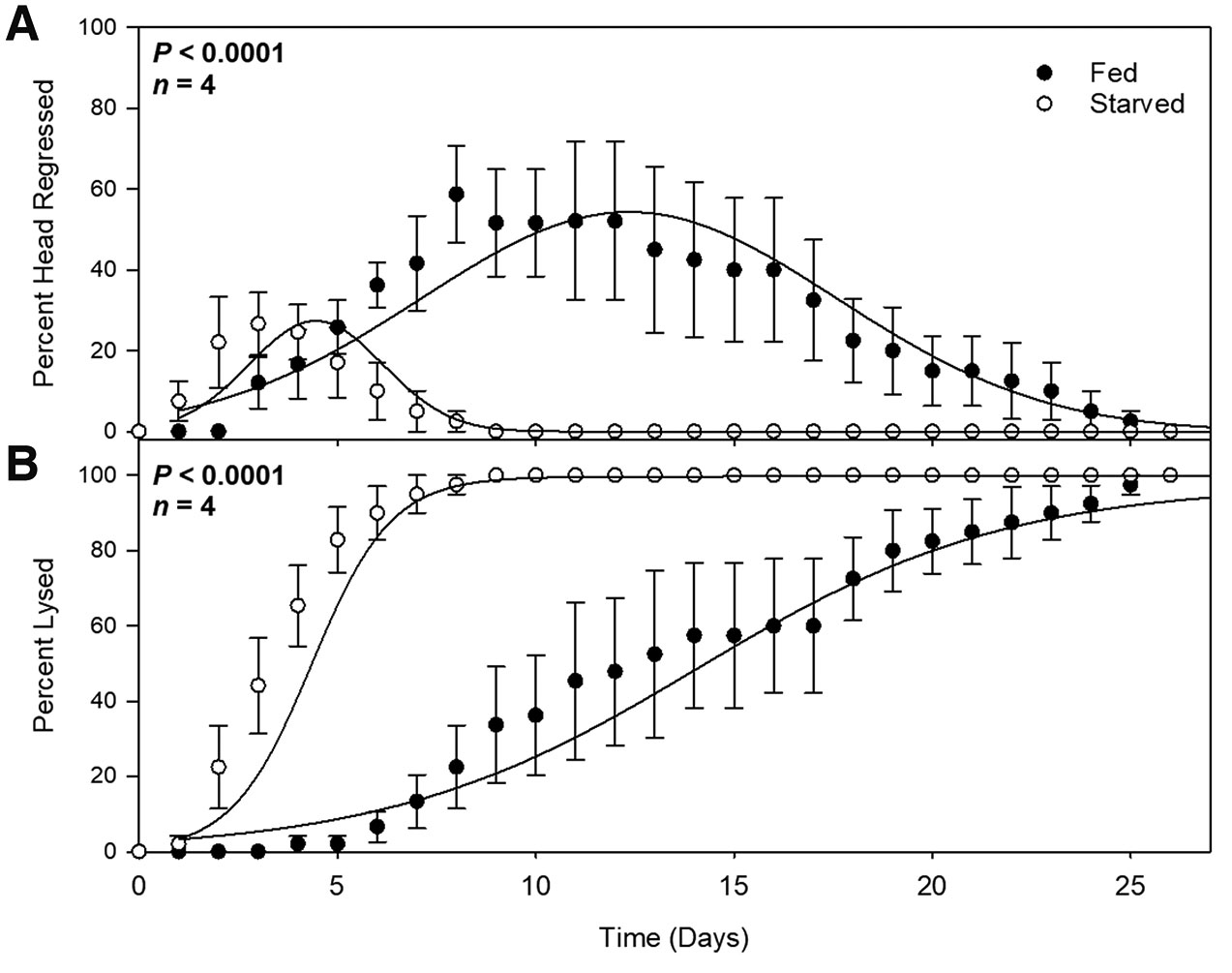
We hypothesized that if autophagy is required for homeostatic tissue maintenance, then chloroquine treatment would lead to phenotypes in whole unamputated planarians. We also assumed that planarians under prolonged starvation would be particularly sensitive to chloroquine treatment because of a possible autophagy requirement in an energy-depleted state. We starved planarians for 40 days to ensure that they had transitioned to degrowth (Fig. 1A). This timeline was chosen because of previous planarian studies showing increased apoptotic cell death after a 40-day starvation period (Pellettieri et al., 2010). A “Fed” group of planarians was then provided pureed calf liver on three alternating days while a “starved” group was not fed during this time (Fig. 1A). Both groups were then treated with chloroquine, and planarians were monitored daily for resulting phenotypes. While we expected to observe animal-wide tissue lysis due to an expected autophagy requirement in all cell types, we surprisingly found that the first phenotype induced by chloroquine was head regression and loss of tissues anterior to the pharynx (Fig. 1B). This is consistent with recent RNAi experiments designed to knockdown planarian atg1 function, where loss of atg1 resulted in head regression (Ma et al., 2025). For both head regression and tissue lysis, we found that the starved group was significantly more sensitive to chloroquine treatment with the starved group showing peak head regression at day 4 and the fed group at day 13 (Fig. 1B and Fig. 2). This is consistent with an increased requirement for autophagy in starved planarians and indicates that these chloroquine-induced effects are energy-dependent and not due to general drug toxicity.
Fig. 1. Chloroquine induces tissue maintenance phenotypes.
(A) Molecular structure of chloroquine. (B) Schedule of starvation period, feeding and chloroquine treatment. Two groups of planarians were starved for 40 days. The “Fed” group was fed on days 40, 42, and 44, while the “Starved” group was not fed on these days. Both groups were treated with 10 µM chloroquine and monitored for tissue maintenance phenotypes every 24 h for the duration of the experiment. (C) Representative images demonstrating the effects of chloroquine treatment on whole unamputated animals. In both “Fed” and “Starved” groups, chloroquine induced head regression (arrows) and subsequent tissue lysis. Scale bars: 500 µm.
Fig. 2. Chloroquine-induced phenotypes occur earlier in starved planarians.
Quantifications of percentage of planarians exhibiting head regression (A) or tissue lysis (B) in “Fed” (closed circles) and “Starved” (open circles) groups over twenty-five days. Each data point represents a daily average. N=4 independent replicates (9-10 animals per treatment, per experiment). Error bars indicate standard error, and probability values were calculated using analysis of variance with repeated measures using a General Linear Mixed Model (PROC GLIMMIX, SAS Studio 3.82).
As head regression is one of the characteristic phenotypes of planarian stem cell loss or dysfunction, this result suggests that autophagy is required for normal neoblast behavior. However, we did not observe ventral curling, another hallmark of neoblast deficiency, instead chloroquine treatment resulted in tissue lysis and death, suggesting a role for autophagy in non-neoblast cells, perhaps in cells that are participating in cell turnover or tissue remodeling. A recent study unveiled that planarian epidermal turnover rate is significantly higher on the ventral side compared to that on the dorsal side (Lee et al., 2024). We imagine that ventral curling phenotypes observed in neoblast-deficient worms are due to the high ventral cell turnover rate without the replenishment of new neoblast-derived cells. The absence of ventral curling observed in our chloroquine-treated animals is likely due to a proposed effect it has on epidermal uptake into phagocytes. Head regression observed in chloroquine-treated planarians might not be a neoblast-specific phenotype but due to the failure to broadly maintain anterior tissues in the absence of autophagy. This fits with studies highlighting atg gene expression in tissues including intestinal branches, epidermis, and cephalic ganglia but primarily outside of the piwi-positive neoblast population (Jin et al., 2022; Ma et al., 2025; Ma et al., 2018). We propose that chloroquine-induced head regression occurs because anterior tissues require a higher rate of autophagic flux for anterior tissue maintenance.
Chloroquine leads to regeneration delay and loss of photoreceptor regeneration
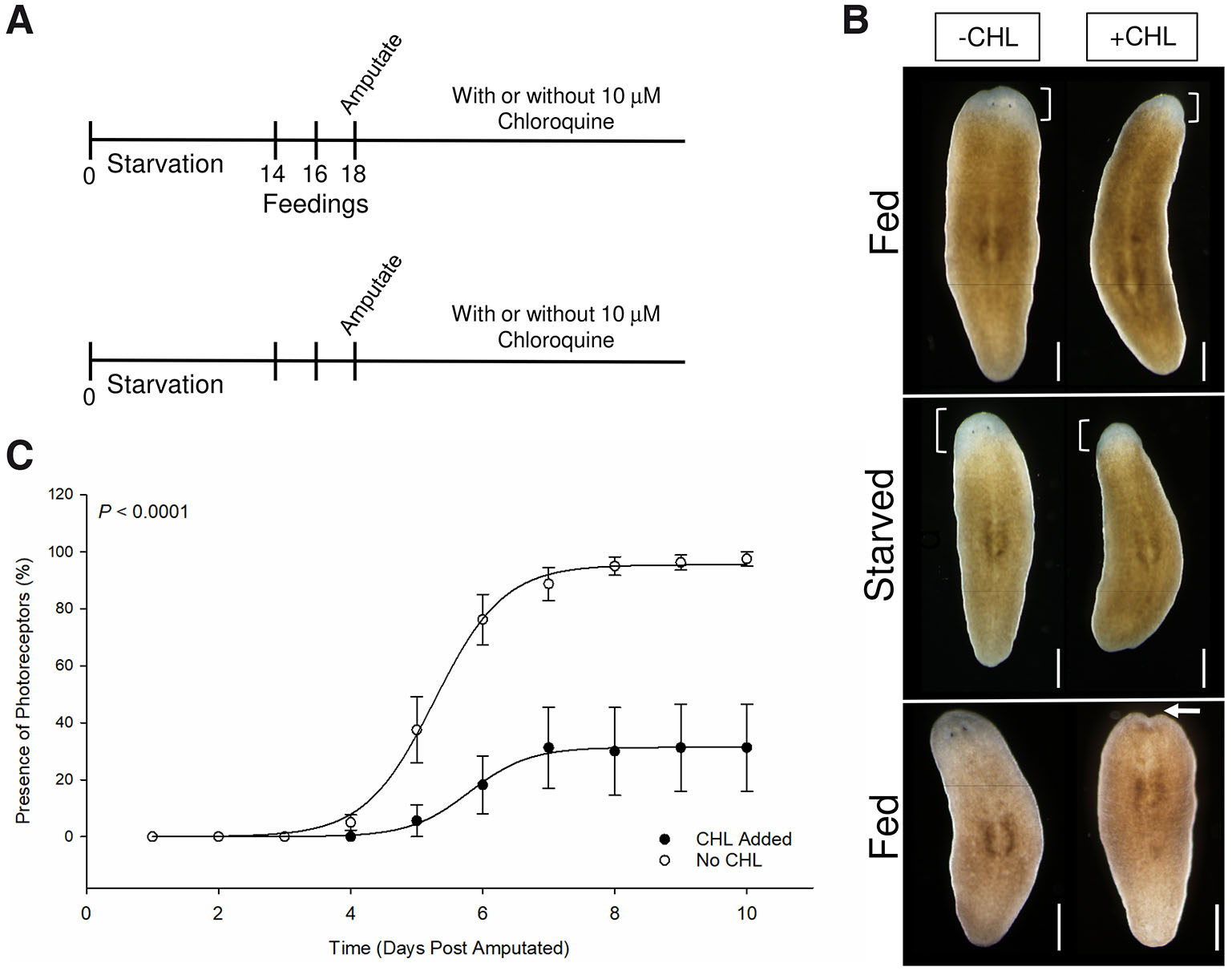
Head regression observed in whole unamputated planarians could be due to neoblast failure to contribute new differentiated cells or due to disruptions in general anterior cell maintenance and turnover. Therefore, we assessed whether chloroquine inhibits the ability for planarians to regenerate lost tissue following amputation. Because blastemal cells are post-mitotic and constructed from neoblast division progeny, we reasoned that failure to form a blastema would indicate chloroquine sensitivity in early neoblast function. We starved planarians for 14 days prior to the start of the experiment (Fig. 3A). A “Fed” group of planarians was then provided pureed calf liver on three alternating days while a “starved” group was not fed during this time (Fig. 3A). Head regions were amputated and posterior tail fragments were isolated and left untreated or treated with chloroquine. Tail fragments were monitored daily for regenerative growth.
Fig. 3. Chloroquine affects photoreceptor regeneration and rate.
(A) Schedule of starvation period, feeding, amputation and chloroquine treatment. Four groups of planarians were starved for 14 days. The “Fed” group was fed on days 14, 16 and 18, while the “Starved” group was not fed on these days. Some groups were treated with 10 µM chloroquine or left untreated. Planarian heads were amputated and posterior fragments were monitored for tissue regeneration phenotypes every 24 h for the duration of the experiment. (B) Representative images at 6 dpa (top two rows) demonstrating the effects of chloroquine treatment on regenerating posterior fragments. In both “Fed” and “Starved” groups, chloroquine did not completely inhibit blastemal formation (brackets), but did negatively impact regeneration speed and photoreceptor regeneration. Fragments in bottom row exhibited head regression by 14dpa following extensive blastemal growth. (C) Comparing the percentage of fragments containing regenerated photoreceptors between chloroquine-treated (closed circles) and untreated (open circles) groups over ten days. Each data point represents a daily average. N=4 independent replicates (9-10 animals per treatment per experiment). Error bars indicate standard error and probability values were calculated using analysis of variance with repeated measures using a General Linear Mixed Model (PROC GLIMMIX, SAS Studio 3.82). Scale bars: 500µm.
We observed that chloroquine did not inhibit regeneration as tail fragments formed an anterior unpigmented blastema in treated groups (Fig. 3B). This is consistent with previous studies where loss of select autophagy-related genes results in morphological defects but does not preclude blastemal growth (Jin et al., 2022; Kang et al., 2019; Ma et al., 2025). We used the reemergence of photoreceptors over time as a measure of regeneration rate and found that chloroquine causes delays in regeneration between days 4 and 7 (Fig. 3C). In addition to this, although we did not directly measure blastema size, we made note of smaller blastemas in the chloroquine-treated group compared to untreated controls. We did not detect a statistically significant difference with chloroquine treatment between starved and fed groups. However, a phenotype observed in chloroquine treated tail fragments is the failure to regenerate photoreceptors in both starved and fed groups (Fig. 3 B,C). While 100% of untreated fragments regenerated photoreceptors by day 7, only 25% of chloroquine-treated fragments regenerated photoreceptors by day 10 (Fig. 3C).
Together, these data suggest that chloroquine negatively affects homeostatic tissue maintenance and the rate of regeneration but does not inhibit blastema formation. Instead, chloroquine appears to disrupt the regeneration of at least some anterior structures. Consistent with a primary role for autophagy in anterior tissue maintenance and turnover, we discovered that chloroquine-treated fragments that regenerated considerable blastemal tissue subsequently underwent head regression around 14dpa (Fig. 3B). Previous studies have shown that planarian pigment cell and epidermal turnover is mediated through their internalization by intestinal phagocytes and that chloroquine treatment affects turnover by inhibiting this internalization event (Lee et al., 2024; Lindsay-Mosher et al., 2024). Therefore, it is possible that the collapse of anterior tissues is due to a failure to undergo normal turnover and maintenance through a similar mechanism. While chloroquine appeared to promote a faster degrowth rate and tissue lysis, no head regression phenotype was reported (Lee et al., 2024). This could be due to differences in the length of initial starvation time and/or concentration of administered chloroquine. Indeed, we did not observe tissue maintenance phenotypes at 1µM chloroquine.
A gradient of Chloroquine sensitivity along the A/P axis
One possible mechanism by which autophagic flux controls anterior tissue maintenance and turnover is through interaction between anterior tissues and the nearby intestinal network (Lee et al., 2024). The rapid collapse of anterior tissues after chloroquine treatment highlights a greater role for autophagic-mediated turnover in anterior tissues than in posterior tissues. Therefore, we became interested in whether cells along the A/P axis exhibit different sensitivities to chloroquine-induced lysis and death. Planarian axial patterning is controlled by positional control genes (PCGs) that emanate primarily from muscle tissue and form morphogen gradients along all body axes (Cutie et al., 2017; Scimone et al., 2017; Scimone et al., 2016). Members of conserved signaling pathways have been shown to control these patterning systems, and upregulation or downregulation of these signals leads to severe patterning defects and failure to control normal neoblast fate (Petersen and Reddien, 2008, 2009). So, it would be reasonable to suggest that levels of autophagy-based turnover would vary along the A/P axis. We hypothesized that if this were the case, then isolated fragments along the A/P axis would exhibit different levels of lysis in response to chloroquine.
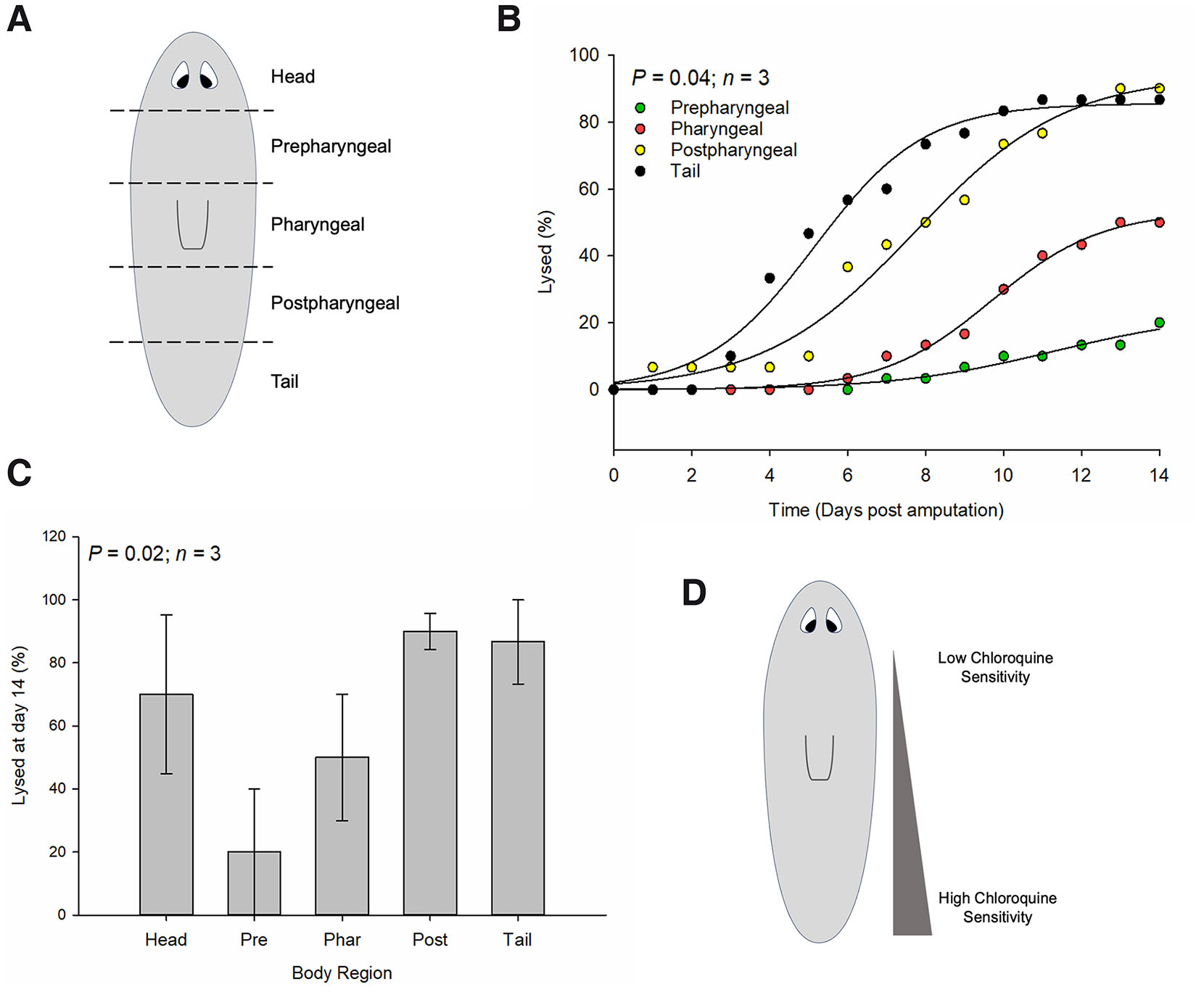
To assess whether chloroquine sensitivity varies across the A/P axis, we starved planarians for 14 days prior to the start of the experiment. Planarians were then amputated at four A/P positions to give rise to head, prepharyngeal, pharyngeal, postpharyngeal, and tail fragments (Fig. 4A). Fragments were then treated with chloroquine or left untreated, and the number of fragments that underwent lysis over time was recorded. We found that fragments underwent lysis at different rates according to their A/P position with lysis first observed in tail fragments (Fig. 4B). Lysis was observed in a tail-to-postpharyngeal-to-pharyngeal-to-prepharyngeal order over time. Prepharyngeal fragments exhibited significantly less lysis in response to chloroquine suggesting that chloroquine sensitivity varies along the planarian A/P axis and that the prepharyngeal region is more resistant to autophagy inhibition (Fig. 4D). Indeed, an average of 20% of prepharyngeal fragments underwent lysis by day 14 (Fig. 4C). This is compared to the number of tail (86.6%), postpharyngeal (90%), and head (70%) fragments that lysed during this time. In our experiment, head fragments appear to lyse independently of this A/P gradient.
Fig. 4. Chloroquine sensitivity across the planarian anterior-posterior (A/P) axis.
(A) Planarians were amputated to give rise to five fragments along the A/P axis (head, prepharyngeal, pharyngeal, postpharyngeal, and tail) and treated with chloroquine. Planarian tissue lysis was monitored daily for two weeks. (B) Quantifications of percentage of planarian fragments (excluding head fragments for clarity) that underwent tissue lysis over time. N=3 independent replicates (9-10 animals per treatment per experiment). (C) Quantifications of percentage of planarian fragments that underwent tissue lysis by day 14. N=3 independent replicates (9-10 animals per treatment per experiment). Error bars indicate standard error and probability values were calculated using analysis of variance with repeated measures using a General Linear Mixed Model (PROC GLIMMIX, SAS Studio 3.82). (D) Representation of a proposed gradient of chloroquine sensitivity in the planarian. Highest sensitivity is observed in tail fragments and tapers off towards prepharyngeal region. Prepharyngeal fragments were highly resistant to chloroquine’s effects on tissue lysis/death.
Prepharyngeal resistance to autophagy inhibition could be due to high endogenous levels or a higher rate of autophagic flux in that region, thus partially masking the effects of chloroquine. Although a more direct observation using markers of autophagy will have to be performed, there are genes that are enriched in prepharyngeal tissues that lend support to this. Expression analyses indicate that the homolog of human sirtuin-1 (Smed-Sirt-1) is expressed broadly in neoblasts and differentiated cells but is enriched in the prepharyngeal region (Ziman et al., 2020). While knockdown of its function leads to a decrease in autophagy markers and growth, it does not disrupt regeneration. Conversely, promotion of sirtuin function with resveratrol treatment increases growth. Another study unveiled the function of a novel family of genes in triclads, called blitzschnell (bls), that are enriched in prepharyngeal secretory cells (Pascual-Carreras et al., 2020). Proposed to act as a nutrient sensor, bls gene expression is induced by feeding and has been shown to balance cell proliferation, apoptosis, and cell size in a nutrient-dependent fashion. Although no autophagy role for bls genes has been reported, they have been shown to interact with the insulin/Akt/mTOR pathway, a well characterized upstream inhibitor of autophagy and regulator of planarian tissue maintenance (González-Estévez et al., 2012; Oviedo et al., 2008; Tu et al., 2012).
We propose a model whereby an A/P gradient of autophagic-based turnover in the planarian exists with the highest level present in the prepharyngeal region. This would allow for preferential maintenance and turnover of anterior tissues in response to nutrient levels or other external stressors. This might explain why anterior tissue regression occurs in neoblast-deficient, chloroquine-treated, and atg1 RNAi-treated planarians. Additionally, a gradient of autophagic turnover rate might explain why some flatworms have lost regenerative capacity in posterior regions but maintain it in the head, and how reversal of A/P patterning signals rescue regenerative potential (Liu et al., 2013; Sikes and Newmark, 2013). Pharmacological approaches that induce autophagy have been shown to improve regeneration in a variety of model systems. Treatment with rapamycin, resveratrol, and metformin, known inhibitors of the mTOR complex and activators of autophagy, improves nerve, muscle, and liver regeneration in rodents and heart regeneration in zebrafish. This indicates that autophagy plays an instructive role in promoting tissue regeneration and maintenance and lends support to the possibility that therapeutically increasing autophagic-based turnover rate might promote regeneration in aging or regeneration-deficient tissues.
Materials and Methods
Planarian care and chloroquine treatment
The CIW4 asexual strain of Schmidtea mediterranea was maintained in 1X Montjuïc salts (1.6mM NaCl, 1.0mM CaCl2, 1.0mM MgSO4, 0.1mM MgCl2, and 0.1mM KCl; pH 7.0) in MilliQ water at 21°C (Cebrià and Newmark, 2005). Planarians were fed pureed calf liver once a week until the start of each experiment discussed below. Chloroquine treatment was conducted using chloroquine diphosphate salt (Sigma-Aldrich) diluted with 1X Montjuïc salts in MilliQ water. Tested concentrations ranged from 1µM to 100µM and were guided by previous studies done in hydra and mice. Each experiment reported in this study was performed using 10µM chloroquine. Microscopic images were obtained on an Olympus SZX10 stereomicroscope and cropped using ImageJ software.
Chloroquine treatment in whole unamputated planarians
For each experiment, 20 planarians were starved for 40 days prior to being separated into two equal numbered groups. One group (“Fed”) was fed pureed calf liver on days 40, 42, and 44 while the other group (“Starved”) was not fed on those days. Both groups were treated with 10µM chloroquine in planarian water starting on day 46. Chloroquine was replaced every 48 hours for the duration of the experiment. We monitored planarians daily for head regression and lysis/death. This experiment was replicated three times.
Chloroquine treatment in amputated planarians
For each experiment, 40 planarians were starved for 14 days prior to being separated into two equal numbered groups. One group (“Fed”) was fed pureed calf liver on days 14, 16, and 18 while the other group (“Starved”) was not fed on those days. On day 21, all planarians were transversely amputated anterior to the pharynx and both “Fed” and “Starved” groups divided into two equal numbered groups (“-Chl” and “+Chl”). While the “-Chl” groups were left untreated, the “+Chl” groups were treated with 10µM chloroquine in planarian water starting on day 21. Chloroquine was replaced every 48 hours for the duration of the experiment. We monitored planarians daily for blastema and photoreceptor regeneration. This experiment was replicated four times.
Assaying chloroquine sensitivity along the anterior-posterior (A/P) axis
For each experiment, 20 planarians were starved for 14 days. On day 14, all planarians were transversely amputated along four amputation planes along the A/P axis, giving rise to head, pre-pharyngeal, pharyngeal, post-pharyngeal and tail fragments. 10 control and 10 chloroquine-treated fragments from each A/P position were placed into their own container. While the control group was left untreated, the chloroquine-treated group was treated with 10 µM chloroquine in planarian water starting on day 14. Chloroquine was replaced every 24 hours for the duration of the experiment. We monitored planarians daily for tissue lysis/death. This experiment was replicated three times.
Statistical analysis
In each experiment (i.e., starvation, amputation, and body region), containers that held planarians received the chloroquine treatment and served as the experimental unit (EU). Individual worms served as subsamples and data were expressed as a percentage using Mean and Standard Error across all trials (Replicates). Analysis of Variance with repeated measures was performed using a General Linear Mixed Model procedure (PROC GLIMMIX) in SAS Studio 3.82 where Day (Time) was the Random Variable. We used an Autoregressive Covariance Matrix (AR(1)) and differences among treatment effects was explored using the PDIFF function within the LSMEANS statement.
Abbreviations
A/P, anterior-posterior ; ATG, autophagy-related gene ; CHL, chloroquine ; DAP-1, death-associated protein-1 ; DPA, days post-amputation ; PCG, positional control gene ;Declarations
Author contributions
R.J. performed the unamputated and tissue regeneration experiments. J.T. performed the A/P axis sensitivity experiments. R.J.M. was primarily responsible for experimental design and writing the manuscript. M.T. helped with experimental design and statistical analysis.
Competing interests
The authors declare no competing or financial interests.
References
Adell T., Cebrià F., Abril J. F., Araújo S. J., Corominas M., Morey M., Serras F., González-Estévez C. (2025). Cell death in regeneration and cell turnover: Lessons from planarians and Drosophila. Seminars in Cell & Developmental Biology 169: 103605.
Adell T., Cebria F., Salo E. (2010). Gradients in Planarian Regeneration and Homeostasis. Cold Spring Harbor Perspectives in Biology 2: a000505-a000505.
Allen E. A., Baehrecke E. H. (2020). Autophagy in animal development. Cell Death & Differentiation 27: 903-918.
Aman Y., Schmauck-Medina T., Hansen M., Morimoto R. I., Simon A. K., Bjedov I., Palikaras K., Simonsen A., Johansen T., Tavernarakis N., Rubinsztein D. C., Partridge L., Kroemer G., Labbadia J., Fang E. F. (2021). Autophagy in healthy aging and disease. Nature Aging 1: 634-650.
Ballabio A., Bonifacino J. S. (2020). Lysosomes as dynamic regulators of cell and organismal homeostasis. Nature Reviews Molecular Cell Biology 21: 101-118.
Balnis J., Jackson E. L., Drake L. A., Singer D. V., Bossardi Ramos R., Singer H. A., Jaitovich A. (2025). Rapamycin improves satellite cells’ autophagy and muscle regeneration during hypercapnia. JCI Insight 10: e182842.
Bossert P., Thomsen G. H. (2017). Inducing Complete Polyp Regeneration from the Aboral Physa of the Starlet Sea Anemone Nematostella vectensis. Journal of Visualized Experiments 54626.
Boya P., Codogno P., Rodriguez-Muela N. (2018). Autophagy in stem cells: repair, remodelling and metabolic reprogramming. Development 145: dev146506.
Cebrià F., Newmark P. A. (2005). Planarian homologs of netrin and netrin receptor are required for proper regeneration of the central nervous system and the maintenance of nervous system architecture. Development 132: 3691-3703.
Chen F., Pu S., Tian L., Zhang H., Zhou H., Yan Y., Hu X., Wu Q., Chen X., Cheng S. H., Xu S. (2024). Radix Rehmanniae Praeparata promoted zebrafish fin regeneration through aryl hydrocarbon receptor-dependent autophagy. Journal of Ethnopharmacology 331: 118272.
Cutie S., Hoang A. T., Payumo A. Y., Huang G. N. (2017). Unconventional Functions of Muscles in Planarian Regeneration. Developmental Cell 43: 657-658.
Ding Z., Cao J., Shen Y., Zou Y., Yang X., Zhou W., Guo Q., Huang C. (2018). Resveratrol Promotes Nerve Regeneration via Activation of p300 Acetyltransferase-Mediated VEGF Signaling in a Rat Model of Sciatic Nerve Crush Injury. Frontiers in Neuroscience 12: 341.
Du J., Liu W., Song Y., Zhang Y., Dong C., Xiong S., Huang Z., Wang T., Ding J., He Q., Yu Z., Ma X. (2024). Activating autophagy promotes skin regeneration induced by mechanical stretch during tissue expansion. Burns & Trauma 12: tkad057.
Eisenhoffer G. T., Kang H., Alvarado A. S. (2008). Molecular Analysis of Stem Cells and Their Descendants during Cell Turnover and Regeneration in the Planarian Schmidtea mediterranea. Cell Stem Cell 3: 327-339.
Emili E., Pérez-Posada A., Vanni V., Salamanca-Díaz D., Ródriguez-Fernández D., Christodoulou M. D., Solana J. (2025). Allometry of cell types in planarians by single-cell transcriptomics. Science Advances 11: eadm7042.
González-Estévez C., Felix D. A., Aboobaker A. A., Saló E. (2007a). Gtdap-1 and the Role of Autophagy During Planarian Regeneration and Starvation. Autophagy 3: 640-642.
González-Estévez C., Felix D. A., Aboobaker A. A., Saló E. (2007b). Gtdap-1 promotes autophagy and is required for planarian remodeling during regeneration and starvation. Proceedings of the National Academy of Sciences 104: 13373-13378.
González-Estévez C., Felix D. A., Smith M. D., Paps J., Morley S. J., James V., Sharp T. V., Aboobaker A. A. (2012). SMG-1 and mTORC1 Act Antagonistically to Regulate Response to Injury and Growth in Planarians. PLoS Genetics 8: e1002619.
Hansen M., Rubinsztein D. C., Walker D. W. (2018). Autophagy as a promoter of longevity: insights from model organisms. Nature Reviews Molecular Cell Biology 19: 579-593.
Huang H., Chen L., Zhang H., Li S., Liu P., Zhao T., Li C. (2016). Autophagy Promotes Peripheral Nerve Regeneration and Motor Recovery Following Sciatic Nerve Crush Injury in Rats. Journal of Molecular Neuroscience 58: 416-423.
Jin B., Ren J., Chen J., Dong Z., Chen G., Liu D. (2022). Autophagy-related DjAtg1-1 plays critical role in planarian regeneration by regulating proliferation and cell death. Cell and Tissue Research 388: 273-286.
Kang J., Dong Z., Wang J., Chen G., Liu D. (2019). Autophagy-related Djatg8 is required for remodeling in planarians Dugesia japonica. Biology Open 8: bio045013.
Lee J.R., Boothe T., Mauksch C., Thommen A., Rink J. C. (2024). Epidermal turnover in the planarian Schmidtea mediterranea involves basal cell extrusion and intestinal digestion. Cell Reports 43: 114305.
Lindsay-Mosher N., Lusk S., Pearson B. J. (2024). Planarians require ced-12/elmo-1 to clear dead cells by excretion through the gut. Cell Reports 43: 113621.
Liu S.Y., Selck C., Friedrich B., Lutz R., Vila-Farré M., Dahl A., Brandl H., Lakshmanaperumal N., Henry I., Rink J. C. (2013). Reactivating head regrowth in a regeneration-deficient planarian species. Nature 500: 81-84.
Ma K., Guo F., Li R., Song G., Zhang H., Lu Q., Ma K., Gong S. (2025). Knockdown of Atg1 Impairs Brain Regeneration and Downregulates ECM-Related Genes in the Planarian Dugesia japonica. Molecular Neurobiology 62: 11330-11347.
Ma K., Zhang Y., Song G., Wu M., Chen G. (2018). Identification of Autophagy-Related Gene 7 and Autophagic Cell Death in the Planarian Dugesia japonica. Frontiers in Physiology 9: 1223.
Mauthe M., Orhon I., Rocchi C., Zhou X., Luhr M., Hijlkema K.J., Coppes R. P., Engedal N., Mari M., Reggiori F. (2018). Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14: 1435-1455.
Nagy P., Sándor G. O., Juhász G. (2018). Autophagy maintains stem cells and intestinal homeostasis in Drosophila. Scientific Reports 8: 4644.
Newmark P. A., Sánchez Alvarado A. (2000). Bromodeoxyuridine Specifically Labels the Regenerative Stem Cells of Planarians. Developmental Biology 220: 142-153.
Nichenko A. S., Southern W. M., Atuan M., Luan J., Peissig K. B., Foltz S. J., Beedle A. M., Warren G. L., Call J. A. (2016). Mitochondrial maintenance via autophagy contributes to functional skeletal muscle regeneration and remodeling. American Journal of Physiology-Cell Physiology 311: C190-C200.
Oviedo N. J., Pearson B. J., Levin M., Sánchez Alvarado A. (2008). Planarian PTEN homologs regulate stem cells and regeneration through TOR signaling. Disease Models & Mechanisms 1: 131-143.
Pascual-Carreras E., Marin-Barba M., Herrera-Úbeda C., Font-Martín D., Eckelt K., de Sousa N., García-Fernández J., Saló E., Adell T. (2020). Planarian cell number depends on blitzschnell , a novel gene family that balances cell proliferation and cell death. Development 147: dev184044.
Pellettieri J. (2019). Regenerative tissue remodeling in planarians – The mysteries of morphallaxis. Seminars in Cell & Developmental Biology 87: 13-21.
Pellettieri J., Fitzgerald P., Watanabe S., Mancuso J., Green D. R., Sánchez Alvarado A. (2010). Cell death and tissue remodeling in planarian regeneration. Developmental Biology 338: 76-85.
Peng Z.L., Yin B.X., Ren R.M., Liao Y.L., Cai H., Wang H. (2021). Altered metabolic state impedes limb regeneration in salamanders. Zoological Research 42: 772-782.
Petersen C. P., Reddien P. W. (2008). Smed- β catenin-1 Is Required for Anteroposterior Blastema Polarity in Planarian Regeneration. Science 319: 327-330.
Petersen C. P., Reddien P. W. (2009). A wound-induced Wnt expression program controls planarian regeneration polarity. Proceedings of the National Academy of Sciences 106: 17061-17066.
Poss K. D., Tanaka E. M. (2024). Hallmarks of regeneration. Cell Stem Cell 31: 1244-1261.
Qiang L., Yang S., Cui Y.H., He Y.Y. (2021). Keratinocyte autophagy enables the activation of keratinocytes and fibroblastsand facilitates wound healing. Autophagy 17: 2128-2143.
Reddien P. W., Alvarado A. S. (2004). Fundamentals of planarian regeneration. Annual Review of Cell and Developmental Biology 20: 725-757.
Saera-Vila A., Kish P. E., Louie K. W., Grzegorski S. J., Klionsky D. J., Kahana A. (2016). Autophagy regulates cytoplasmic remodeling during cell reprogramming in a zebrafish model of muscle regeneration. Autophagy 12: 1864-1875.
Sahu S., Sridhar D., Abnave P., Kosaka N., Dattani A., Thompson J. M., Hill M. A., Aboobaker A. (2021). Ongoing repair of migration-coupled DNA damage allows planarian adult stem cells to reach wound sites. eLife 10: e63779.
Scimone M. L., Cote L. E., Reddien P. W. (2017). Orthogonal muscle fibres have different instructive roles in planarian regeneration. Nature 551: 623-628.
Scimone M. L., Cote L. E., Rogers T., Reddien P. W. (2016). Two FGFRL-Wnt circuits organize the planarian anteroposterior axis. eLife 5: 27063937.
Sikes J. M., Newmark P. A. (2013). Restoration of anterior regeneration in a planarian with limited regenerative ability. Nature 500: 77-80.
Suknovic N., Tomczyk S., Colevret D., Perruchoud C., Galliot B. (2021). The ULK1 kinase, a necessary component of the pro-regenerative and anti-aging machinery in Hydra. Mechanisms of Ageing and Development 194: 111414.
Takeda H., Nishimura K., Agata K. (2009). Planarians Maintain a Constant Ratio of Different Cell Types During Changes in Body Size by Using the Stem Cell System. Zoological Science 26: 805-813.
Tang A. H., Rando T. A. (2014). Induction of autophagy supports the bioenergetic demands of quiescent muscle stem cell activation. The EMBO Journal 33: 2782-2797.
Thommen A., Werner S., Frank O., Philipp J., Knittelfelder O., Quek Y., Fahmy K., Shevchenko A., Friedrich B. M., Jülicher F., Rink J. C. (2019). Body size-dependent energy storage causes Kleiber’s law scaling of the metabolic rate in planarians. eLife 8: e38187.
Tu K. C., Pearson B. J., Sánchez Alvarado A. (2012). TORC1 is required to balance cell proliferation and cell death in planarians. Developmental Biology 365: 458-469.
Varga M., Sass M., Papp D., Takács-Vellai K., Kobolak J., Dinnyés A., Klionsky D. J., Vellai T. (2014). Autophagy is required for zebrafish caudal fin regeneration. Cell Death & Differentiation 21: 547-556.
Wagner D. E., Wang I. E., Reddien P. W. (2011). Clonogenic Neoblasts Are Pluripotent Adult Stem Cells That Underlie Planarian Regeneration. Science 332: 811-816.
Wenemoser D., Reddien P. W. (2010). Planarian regeneration involves distinct stem cell responses to wounds and tissue absence. Developmental Biology 344: 979-991.
Witchley J. N., Mayer M., Wagner D. E., Owen J. H., Reddien P. W. (2013). Muscle Cells Provide Instructions for Planarian Regeneration. Cell Reports 4: 633-641.
Xiang Z., Zhang S., Yao X., Xu L., Hu J., Yin C., Chen J., Xu H. (2021). Resveratrol promotes axonal regeneration after spinal cord injury through activating Wnt/β-catenin signaling pathway. Aging 13: 23603-23619.
Xie F., Xu S., Lu Y., Wong K. F., Sun L., Hasan K. M. M., Ma A. C. H., Tse G., Manno S. H. C., Tian L., Yue J., Cheng S. H. (2021). Metformin accelerates zebrafish heart regeneration by inducing autophagy. npj Regenerative Medicine 6: 62.
Yamamoto H., Zhang S., Mizushima N. (2023). Autophagy genes in biology and disease. Nature Reviews Genetics 24: 382-400.
Ziman B., Karabinis P., Barghouth P., Oviedo N. J. (2020). Sirtuin-1 regulates organismal growth by altering feeding behavior and intestinal morphology in planarians. Journal of Cell Science 133: jcs239467.