Int. J. Dev. Biol. 66: 223 - 233 (2022)
Special Issue: Developmental Biology in Greece
Expression of Hey marks a subset of enteroendocrine cells in the Drosophila embryonic and larval midgut
Original Article | Published: 22 November 2021
Abstract
Hey is a conserved transcription factor of the bHLH-Orange family that participates in the response to Notch signaling in certain tissues. Whereas three Hey paralogues exist in mammalian genomes, Drosophila possesses a single Hey gene. Fly Hey is expressed in the subset of newborn neurons that receive a Notch signal to differentiate them from their sibling cells after the asymmetric division of precursors called ganglion-mother-cells. We used a polyclonal anti-Hey serum and a GFP-tagged transgenic duplication of the Hey locus to examine its expression in tissues outside the nervous system in embryos and larvae. We detected robust Hey expression in the embryonic midgut primordium at the time of birth of enteroendocrine cells, identified by expression of Prospero. Approximately half of the Pros-positive cells were also Hey positive at mid-embryogenesis. By the end of embryogenesis, most enteroendocrine cells had downregulated Hey expression, although it was still detectable at low levels after hatching. Low levels of Hey were also detected in subsets of the epithelial enterocytes at different times. Embryo enteroendocrine Hey expression was found to be Notch dependent. In late third-instar larvae, when few new enteroendocrine cells are born, novel Hey expression was detected in one cell of each sibling pair. In conclusion, Hey is strongly expressed in one of each pair of newly-born enteroendocrine cells. This is consistent with a hypothesis that embryonic enteroendocrine cells are born by an asymmetric division of a precursor, where Notch/Hey probably distinguish between the subtypes of these cells upon their differentiation.
Keywords
Hey bHLH-O, enteroendocrine, midgut, Drosophila, Notch
Introduction
Hey proteins are members of the bHLH-O protein group, a subfamily of bHLH transcription factors distinguished by an ‘Orange’ protein-protein interaction domain. They are closely related to Hes [Hairy/E(spl)] group and named after it as Hairy/E(spl)-like with a Y, due to the tyrosine residue in their C-terminal YRPW motif, substituting tryptophan in the corresponding WRPW motif of Hes. Hey transcription factors, similarly to E(spl), are integral members of the Notch signaling pathway, although they also integrate signal for multiple pathways (BMP/TGF-β, JAK-STAT, RAS and HIF signaling) (Fischer and Gessler, 2007). Until recently, their function was thought to be exclusively repressive (Heisig et al., 2012), but new in vitro studies involving Drosophila Hey support a dual transcriptional activity for this factor as potential activator and repressor in an enhancer context- related manner (Stampfel et al., 2015). Accordingly, a more recent study supports a role of Drosophila Hey in regulating histone tail modifications associated with gene activation in vivo (Flint Brodsly et al., 2019).
Hey transcription factors are involved in critical biological processes such as embryonic development, differentiation processes and mature tissue homeostasis, and their action ranges from binary cell fate decisions to inductive processes and control of proliferation (Weber et al., 2014). The three mammalian Hey proteins (Hey1, Hey2, and HeyL) have been identified as crucial factors in cardiac and vascular development, as well as regulators of neural, muscular and bone development (Weber et al., 2014). In Drosophila, few studies have been conducted on the physiological role of the single orthologue of mammalian Hey proteins (Lu et al. 2008; Monastirioti et al. 2010; Flint Brodsly et al. 2019). Strong Hey expression is observed in the developing central nervous system (CNS) of Drosophila, where it is primarily detected in newly born post-mitotic neurons and a few glia (Monastirioti et al., 2010). During both neurogenic phases of the fly life, in embryonic and in larval stages, neurons are produced from the ganglion mother cell (GMC) precursors that have been generated through asymmetric divisions of the neural stem cells, the Neuroblasts (NBs) (Hartenstein and Wodarz, 2013). In most NB lineages, the GMC precursors produce type “A” and type “B” neurons, or, less often, a neuron and a glia via an asymmetric division regulated by Notch signaling (Skeath and Doe 1998; Lee 2017). During this process, Hey is expressed as a target of the activated Notch pathway in one of the two siblings where Hey determines and establishes the “type A” neuronal fate (Monastirioti et al., 2010). On the other hand, Notch-independent expression of Hey is evident in a small number of neuronal lineages including the larval Mushroom Body (MB) lineages, whose GMCs perform symmetric divisions to produce two identical neurons (Monastirioti et al. 2010; Lee 2017). Besides developing CNS, a recent study has suggested that Hey is an important factor for proper tissue integrity and homeostasis of the adult Drosophila midgut and, consequently, fly survival (Flint Brodsly et al., 2019). It was reported that in adult midgut, Hey functions as an identity guardian of the fully differentiated enterocytes (ECs) constantly regulating the expression of nuclear lamins. According to the data and model presented, Hey regulates EC identity partially by repressing the stem cell related type-B lamin (LamDmO) and by promoting the expression of EC genes such as the homeobox transcription factor Pdm1 and the type-A lamin (LamC), which is necessary to prevent the expression of progenitor genes (Flint Brodsly et al., 2019). Interestingly, this adult expression of Hey is very weak and independent of Notch signaling (Dutta et al. 2015; Flint Brodsly et al. 2019).
In this study, we report on Drosophila Hey expression in the intestinal tissue primordium during embryonic and larval life. In the embryo, robust expression of Hey is related to a subpopulation of the presumptive enteroendocrine (EE) cells which are characterized by the expression of the pro-differentiation factor Prospero (Choksi et al., 2006), while weak expression is observed in a few large nuclei characteristic of the enterocyte cell population. Hey expression in embryonic EE cells is dependent on Notch signaling. In the larva tissue, Hey expression is very weak and associates primarily with a subset of the EE cells, as well as ECs of the anterior midgut. At late larval stages, Hey is re-expressed more strongly in one of the two Pros+ cells that appear de novo within AMP islands (Takashima et al., 2011). Our study identifies another instance of Notch dependent expression of Hey in an endodermally derived tissue, the developing midgut primordia, where it relates to postmitotic EE cells. We propose that Hey functions in an asymmetric division of EE cell precursors that putatively generates different peptidergic EE fates.
Results
Hey expression in the developing midgut during embryogenesis
We have previously reported (Monastirioti et al., 2010) that Drosophila Hey is robustly expressed during embryonic development and the major site of its expression is in the central nervous system (CNS). During the course of these early studies, we had also noticed a scattered pattern of Hey expressing cells when focusing on central domains of the developing embryos. These cells become evident after the onset of Hey expression in the CNS, which takes place at embryonic stage 11 (7h AEL), and they reside in regions dorsal to the ventral nerve cord and posterior to the developing brain lobes (Fig. 1A). In this deep internal areas, tissues of mesodermal (visceral musculature) and endodermal (midgut epithelia) origin form and develop. Therefore, we decided to take a closer look at the observed expression pattern of Hey and ascertain whether it coincides with these non-neuronal developing primordia.
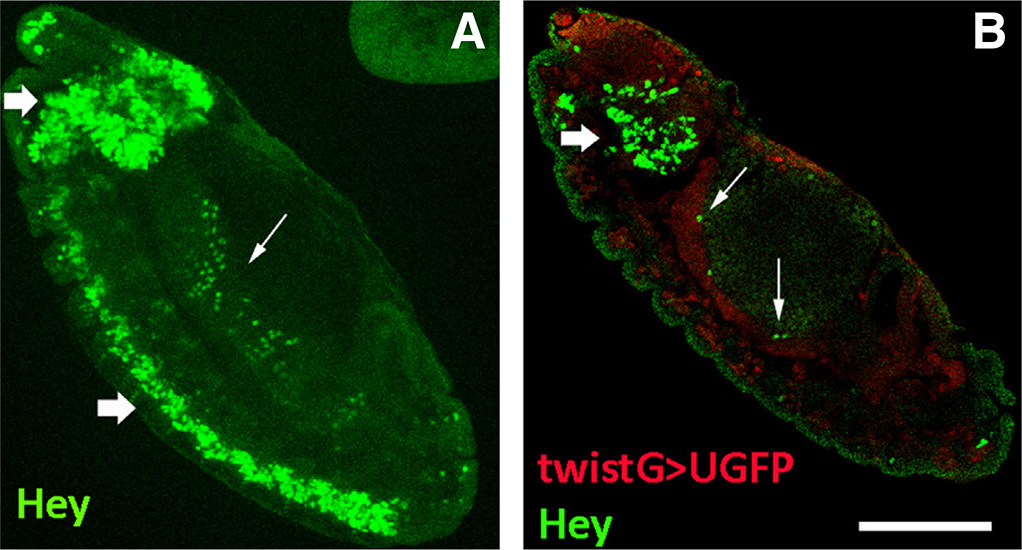
Fig. 1. Hey expression in the Drosophila embryo.
Stage 13 embryos stained for Hey and the mesodermal marker Twist. Anterior is top and ventral is left. (A) Hey (green) is expressed in neurons of the developing CNS (thick arrows) as well as in deep internal areas (arrow). (B) Hey+ cells (green) are distinct from twistGAL4>UAS-GFP positive mesoderm (red) and are localized apically (internally) to the latter (arrows) where midgut primordium develops. Thick arrow points to Hey-positive cells of the developing brain lobe. Scale bar, 40μm.
Table 1
Cell populations in Drosophila embryonic midgut primordia expressing different combinations of transcription factors along with Hey
| Stages 10-11 | stages 12-13 | stage 15 | stage 16 |
| esgGFP+ | esgGFP+ | esgGFP+ | esgGFP+ |
| esgGFP+ pros+ | esgGFP+ pros+ | pros+ | pros+ |
| esgGFP+ Hey+ | esgGFP+ pros+ Hey+ | pros+ Hey+ | pros+ Hey+ |
| pros+ | pdm-1+ | pdm-1+ | |
| pros+ Hey+ | pdm-1+ Hey+ | ||
| pdm-1+ |
Morphogenesis of the Drosophila larval midgut takes place during embryonic life and relies on proliferating endodermal cell clusters and underlying mesodermal cells that will give rise to the functional midgut of the hatching larva and the muscular epithelium that surrounds it, respectively (Hartenstein et al. 1992; Tepass and Hartenstein 1994; Takashima and Hartenstein 2012). The midgut primordium comes from endoderm that invaginates at two sites in the early gastrulating embryo (stage 7) - the anterior and posterior midgut (amg and pmg) invaginations - and is visible at embryonic stages 10-11 as two internalized masses of dividing progenitors at anterior and posterior regions of extended germ band embryos. Initially, these cells undergo an epithelial to mesenchymal transition (EMT) and start migrating towards the middle of the gastrulating embryo, relying on their contact with underlying visceral muscle primordia that develop from mesoderm as well as factors secreted by both themselves and the mesodermal cells (Tepass and Hartenstein 1994; Campbell and Casanova 2015; Pitsidianaki et al. 2021). The majority of cells from the outer layer of the migrating amg and pmg primordia gradually undergo a mesenchymal to epithelial transition (MET); they become postmitotic and give rise to the principal midgut epithelial cells (PMECs) that merge and reorganize into an epithelial layer. Eventually, these cells differentiate into polyploid enterocytes (ECs) that will line the larval midgut epithelium and will be marked by the expression of Pdm1 (Nubbin) transcription factor. A smaller fraction of cells of the inner endodermal masses that stay mesenchymal through migration includes cells with small nuclei that are the progenitors of the adult midgut precursors (AMPs) and of the enteroendocrine (EE) cells of larval midgut. In the pmg, a few cells are distinguished by their large nuclei and correspond to the interstitial cell precursors (ICPs) (Takashima et al. 2011; Takashima and Hartenstein 2012) that will give rise to characteristic enterocytes (interstitial/copper cells) of the acidic middle region of larva midgut (Dubreuil et al. 1998; Overend et al. 2016). Until stage 12/13, all cells of the amg and pmg are labeled by Escargot (Esg), while the precursors of EE cells (pre-EE) are labeled by Prospero (Pros) markers as well. At stage 13, the two rudiments meet in the middle of the embryo, with the ICPs taking position at the amg-pmg fusion site, while by stage 15, the midgut epithelium has formed a closed sac. At this stage, the midgut cell populations have been separated into the undifferentiated AMPs that are the only cells retaining expression of Escargot (Esg+ cells), and two types of postmitotic cells: those that follow the enteroendocrine fate expressing Prospero (Pros+ cells), and the vast majority of the PMECs that make the midgut epithelia and express Pdm1. At this stage, the AMPs and the EEs take positions on the apical site of the PMECs towards the lumen. By stage 16, three constrictions induced by the visceral mesoderm and the expression domains of 4 homeotic genes (Antp, Tsh, Ubx, AbdA) divide the closed epithelium into four domains (Nakagoshi, 2005) that will subsequently develop into the midgut tube of the larval intestine.
Given the observed expression of Hey in embryo regions where midgut and visceral mesoderm tissues develop, our first aim was to further elucidate the tissue and cells that comprise its expression pattern. Initially, we ascertained that Hey expressing cells are distinct from and apical to cells with twistGAL4>UAS-GFP expression characteristic of mesodermal tissues (Fig. 1B). This suggested that Hey+ cells may be part of the developing midgut epithelia, so we performed Hey immunostainings in embryos of different developmental stages in combination with antibodies against protein markers with expression in the different midgut cell populations. The SNAIL family transcription factor Escargot (Esg) characterizes the undifferentiated cells /AMPs (Jiang and Edgar, 2009), the pro-differentiation homeodomain transcription factor Prospero (Pros) is related to EEs, while the polyploid ECs are characterized by the expression of Pdm1 (Nubbin) protein (Micchelli and Perrimon, 2006).
Stages 10-11
At stages 10-11, all cells of both amg and pmg rudiments are expressing Escargot, while Prospero expression is initially observed in Esg+ cells of pmg and shortly afterwards in amg cells too (Takashima et al., 2011; Fig. 2 st11A,st11B). There is intense proliferation activity at these stages (Takashima et al., 2011) among the Esg+ Pros+ cells of both rudiments, as judged by aPH3 (Histone H3-phospho-S10) staining which marks mitotic chromatin (Fig. 4 A-A’’). While robust expression of Hey is already evident in newborn neuronal cells of the extended ventral nerve cord, only weak Hey expression was observed in 3-4 large nuclei at the posterior midgut rudiment of stage 11 embryos (Fig. 2 B-B’’). Although hardly detectable, Hey expression was reproducibly observed in these Esg+ Pros- cells, which we assume correspond to the ICPs, given their large nuclei.
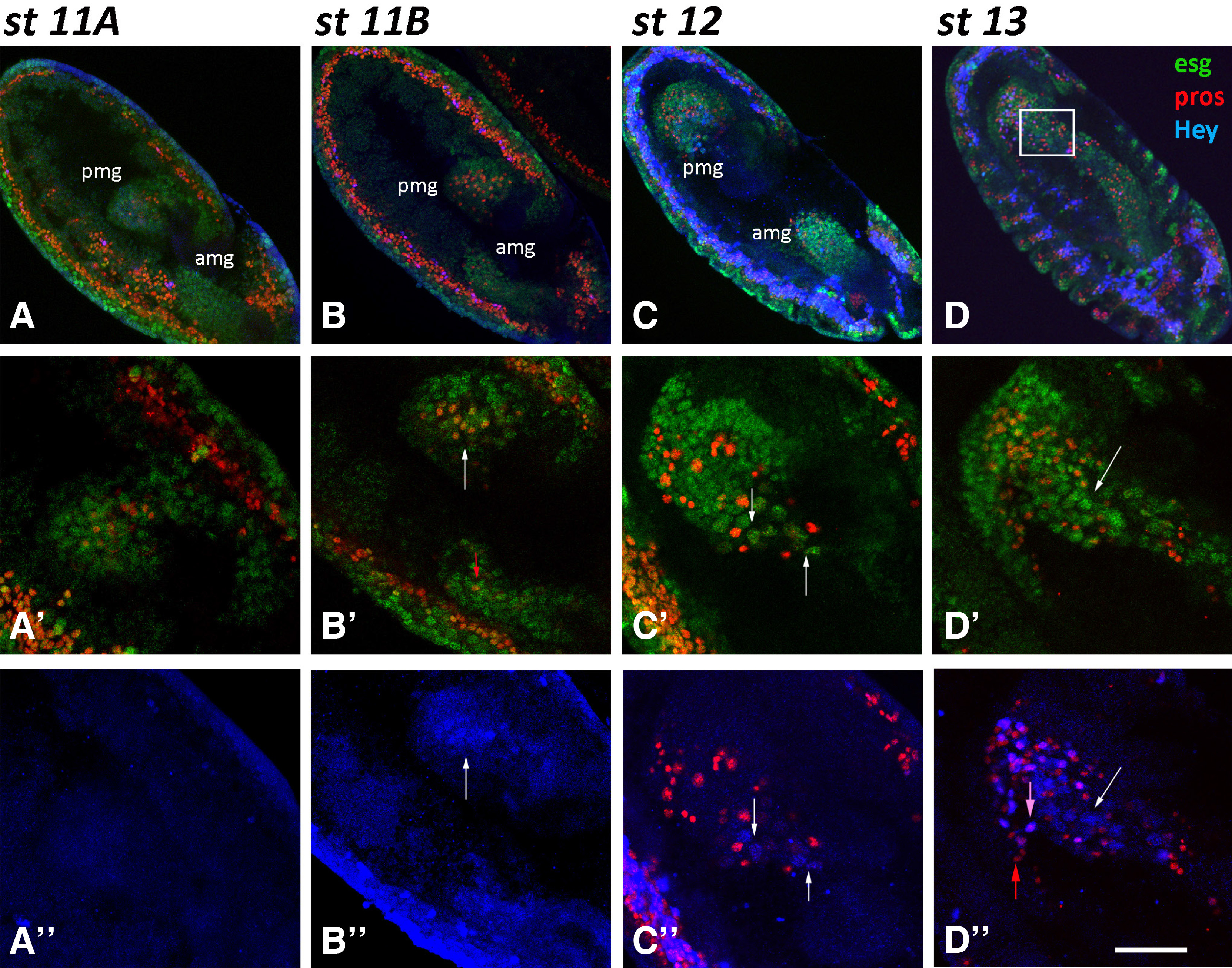
Fig. 2. Hey expression in the midgut primordia during embryogenesis.
Embryos of different stages, as designated, are stained for Esg-GFP (green), Pros (red) and Hey (blue). Anterior is down and ventral is left. (A-A’’, Β-B’’) Two-stage 11-germ band extended embryos of slightly different age in which amg and pmg have invaginated in anterior and posterior ends, respectively. All undifferentiated cells are labeled by Esg-GFP (green). Prospero expression (red) is evident in a subset of the latter, initially in the pmg (A’- st11A) and then in the amg (B’-st11B, small red arrow). Weak Hey expression (blue) is detected in few large nuclei in pmg of the latest st11B embryo (B’, B’’-white arrows). (C-C’’) In stage 12 embryo, amg and pmg primordia are migrating towards each other (C) and large Esg (green), Hey (blue) double positive large nuclei are placed in the front of the moving cell cluster (C’, C’’- white arrows). (D-D’’) In stage 13 embryo, the large Esg (green), Hey (blue) double positive large nuclei (white arrows) are at the fusion point of the amg and pmg primordia. At this stage, robust Hey expression (blue) is also observed in small Pros-positive nuclei (red), but only a subset of the latter (D’’-pink arrow vs red arrow). Scale bar, 40μm (A-D), 23μm (B’-B”, D’-D”), 16μm (A’-A”, C’-C’’).
Stages 12-13
During stages 12 and 13 (7.5-10.5 h AEL), amg and pmg cell populations migrate towards each other following germ-band shortening (Fig. 2 C,st12), and by the end of stage 13 have fused in the middle region (Fig. 2 D,st13). Weak Hey expression is now observed in more Esg+ large nuclei, which are located at the front of the migrating pmg cell cluster (Fig. 2 C’-C’’,st12), while at stage 13 they are found in the middle of the embryo at the point where amg and pmg primordia meet (Fig. 2 D’-D’’, white arrows). The positions occupied by the the large Esg+ weak-Hey+ nuclei during primordia migration further indicate that are putatively a subset of the ICPs.
At this developmental age, besides presumptive ICPs, a few small nuclei initiate Hey expression, initially observed in the posterior midgut primordium and soon afterwards in the anterior one as they still migrate towards each other around late stage 12. Unlike the large ICPs, these small nuclei exhibit robust Hey expression, are all Pros+ (Fig. 2 D’’-pink arrow) and the vast majority of them are also labeled by esg-GFP fusion expression (Fig. 3 A-A’’, arrowheads). Notably, all remaining cells of the primordia continue to express esg-GFP, indicating that they are not fully differentiated yet. The above observations suggest that Hey expressing cells are among the progeny of an EE precursor population that are routed towards acquiring enteroendocrine fate and will soon switch off Escargot expression.
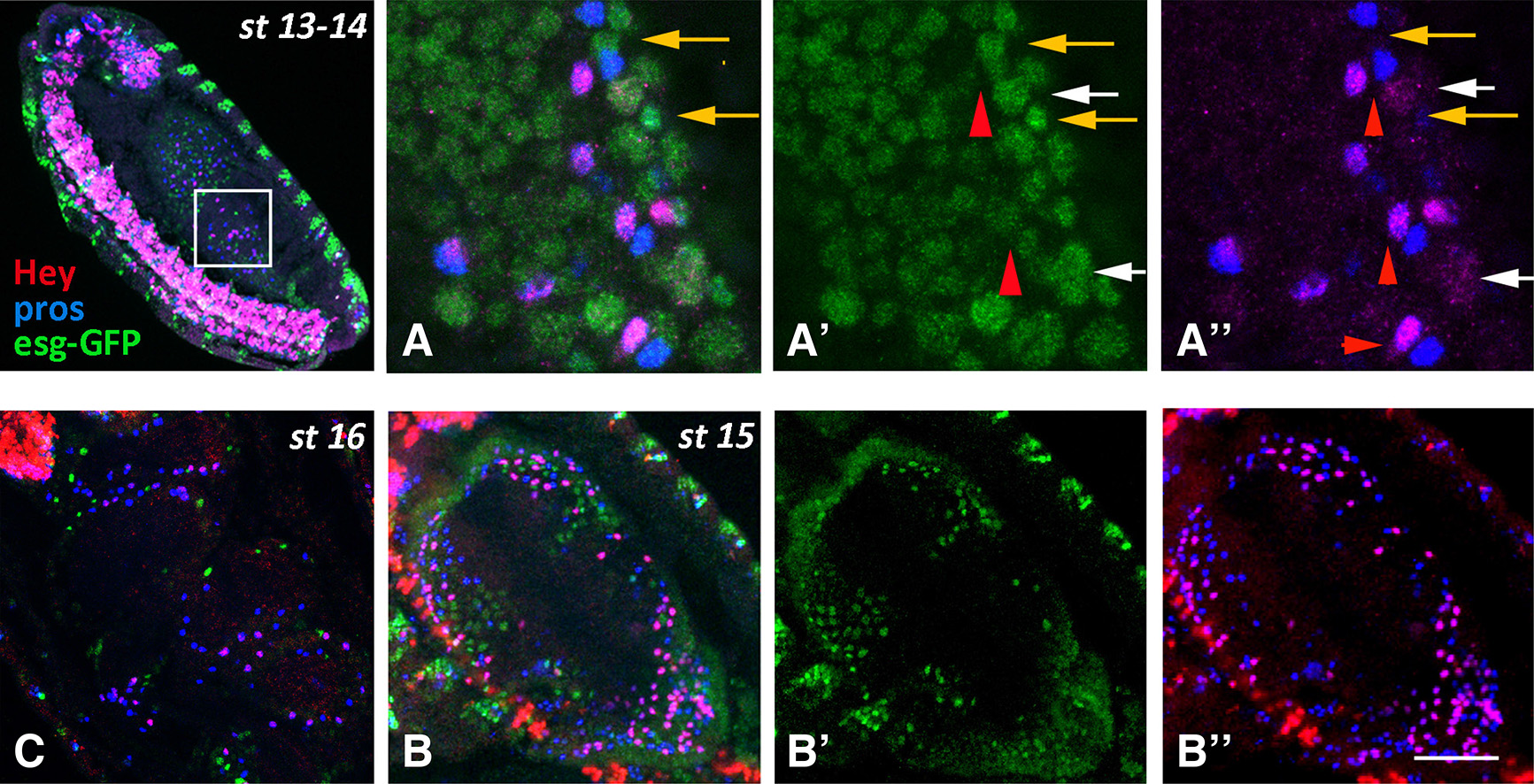
Fig. 3. Hey expression in the midgut primordia during embryogenesis (continued)
Embryos of different stages, as designated, are stained for Esg-GFP (green), Pros (blue) and Hey (red). Anterior is at the top and ventral is to the left. (A-A’’) High magnification images of the boxed area in st13-14 embryo. All cells are still Esg-GFP positive, representing either undifferentiated cells and/or AMPs (A’, A’’-orange arrows). Hey (red) is expressed in a subset of Pros (blue) cells that in some cases exhibit low Esg-GFP staining (A’- red arrowhead). Note that the Pros+Hey+ and Pros+Hey- appear as pairs (A’’- red arrowheads). White arrows in A’-A’’ designate large Esg-GFP+ nuclei with weak Hey staining. (B-B’’) Middle inner area of a stage 15 embryo displaying the lumen of the midgut sac surrounded by the epithelial cells. Esg-GFP+ AMPs (green) and Pros+ and Pros+Hey+ enteroendocrine cells form three distinct populations. Anterior is at the top. (C) Dorsal view of stage 16 embryo at the level of the midgut lumen. At this stage, three constrictions induced by the surrounding mesoderm have divided the midgut sac into four domains. As in stage 15, the Esg-GFP+ AMPs (green) and the Pros+ and Pros+Hey+ enteroendocrine cells form three distinct populations. Scale bar, 6μm (A-A”), 36μm (B-B”), 26μm (C).
During these stages, PH3 immunoreactivity gradually becomes sparser, suggesting that mitotic activity in the midgut primordium is ceasing. The sparse mitotic cells are mostly Pros+ and never Hey+ (Fig. 4 B,C). Actually, Hey expression is found mostly in one of two adjacent Pros+ cells, which may suggest an asymmetric division of an EE progenitor, similarly to what has been observed in the CNS, where Hey is expressed in one of two sibling neurons after a ganglion-mother-cell asymmetric division (Monastirioti et al., 2010).
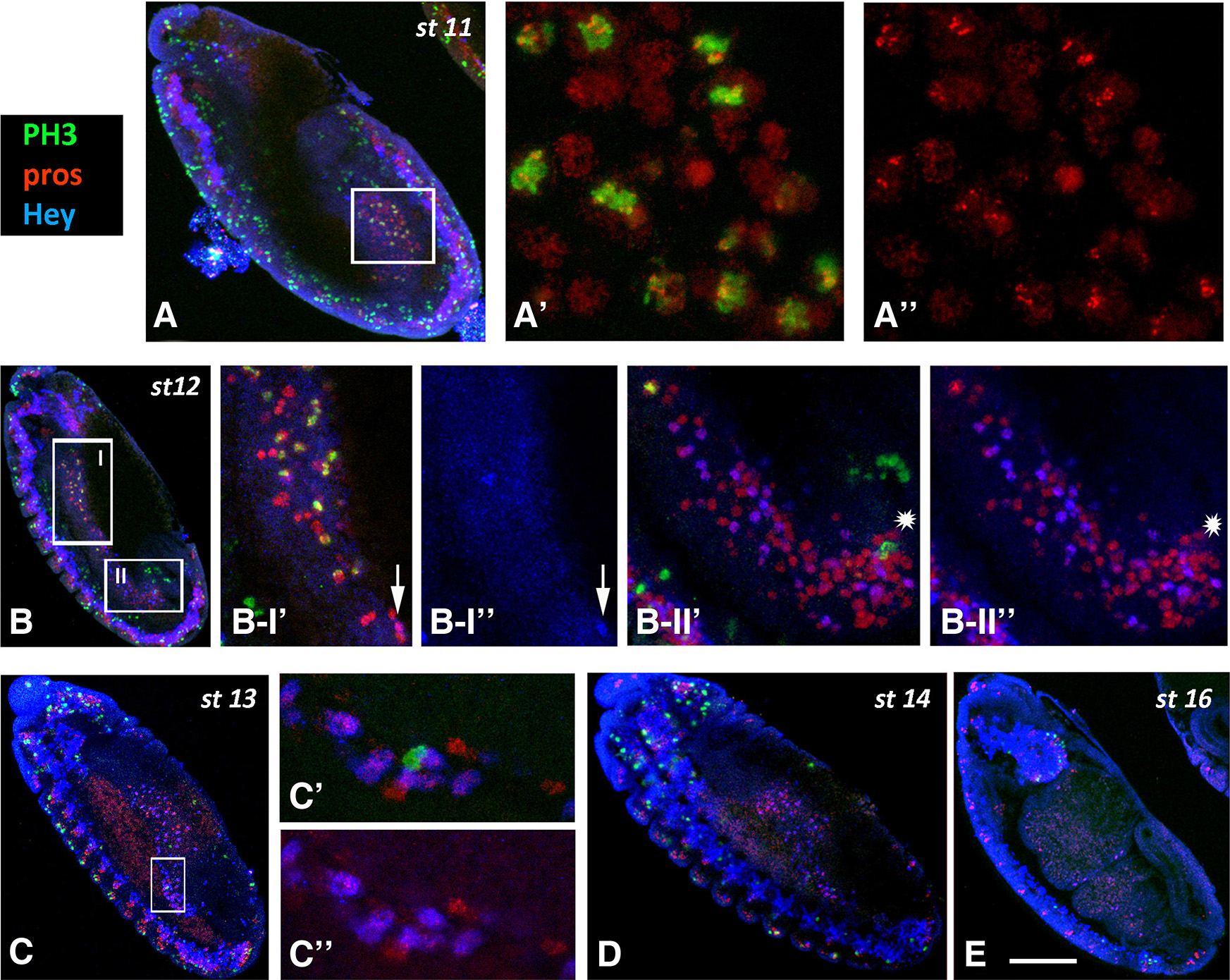
Fig. 4. PH3 immunoreactivity in the midgut primordia during embryogenesis.
Embryos of different stages, as designated, are stained for PH3 (green), Pros (red) and Hey (blue). Anterior is at the top and ventral is to the left. (A-A’’) High magnification images of the boxed area in stage 11 embryo (A). Many Pros+ cells display mitotic activity in the pmg primordium. (B) In stage 12, divisions are still taking place in the amg (B-I) and pmg (B-II) in Pros+ (B-I’) and Pros- (asterisk in B II’-II’’) cells. Although Hey (blue) is already expressed in Pros+ cells we never detected, together with PH3 (arrow in B-I’’). (C-E) In embryos of stage 13 and onwards, divisions in the midgut primordia are sparse (C’, C’’) or missing (D, E). Scale bar, 40μm (A-E), 18μm (B I’-B II”), 6μm (Α’-Α”, C’-C”).
Stages 14-16
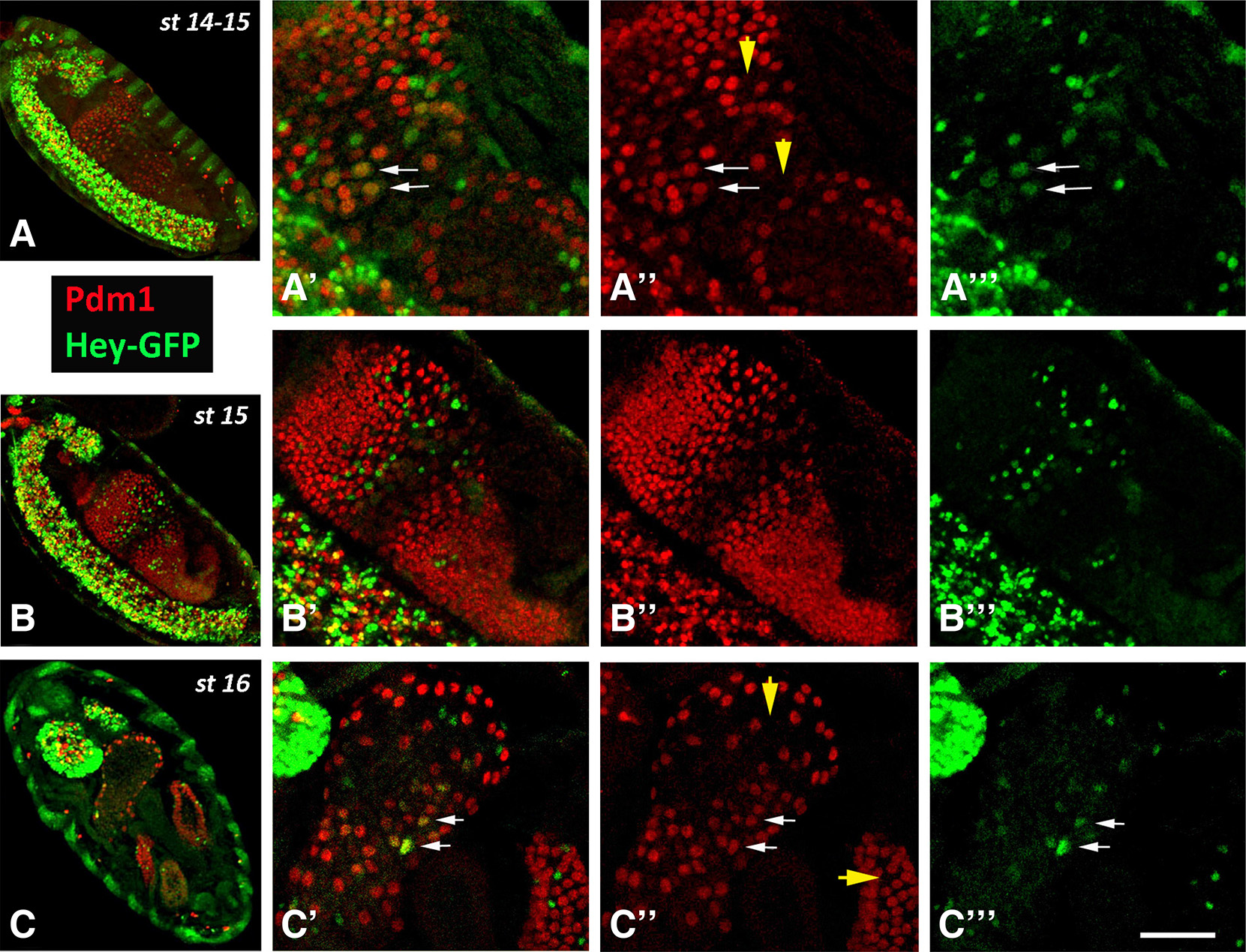
During stages 12-13, when residual mitoses continue in the midgut primordia, the number of Hey expressing cells is rapidly increased, and by stages 14-15 represent a significant subpopulation (almost half the number) of Pros+ EE cells that are devoid of esg-GFP expression. Notably, at stage 15, when the midgut primordium has developed into a closed sac, the different cell types of the epithelium have been established as Esg+ undifferentiated AMPs, Pros+ Hey- and Pros+ Hey+ EE cells (Fig. 3 B-B’’) located towards the lumen apically to Pdm1+ ECs (Fig. S1). At this stage, the large nuclei of the latter class, even the large ICPs, are devoid of Hey expression, although the more stable Hey-GFP fusion (see larva section) is still weakly detectable in the presumptive ICPs (Fig. 5 A-A’’’). Later on, by stage 16, Hey-GFP is detectable in several large Pdm1+ nuclei in the anterior most midgut domain (Fig. 5 C-C’’’), which were occasionally detected by the anti-Hey antibody (data not shown), representing an unknown subpopulation of enterocytes. Strong Hey expression is still primarily detected in Pros+ EEs, albeit in significantly smaller numbers (Fig. 3C, Fig. 5 C-C’’’, Fig. S1 B-B’). Indeed, when we counted all three small nuclei populations, the AMPs (Esg-GFP+) and both types of EEs (Pros+ Hey- and Pros+ Hey+) in stages 15 and 16, we found that the total number of AMPs and EEs remains approximately stable (50 vs 46 AMPs and 185 vs 186 EEs in stage 15 and stage 16, respectively). However, the Pros+ Hey+ EE subpopulation has been significantly reduced (99 vs 69 in stages 15 and 16, respectively) which indicates that Hey expression may be transient during development of midgut primordium and may only be present during establishment of EE cell fate.
Fig. 5. Expression of Hey-GFP in ECs during late embryonic stages.
Late stage embryos of Hey-GFP line stained for GFP (green) and Pdm1 (red). (A-A’’’, B-B’’’) Sagittal views of stage 14-15 embryos. Anterior is at the top and ventral to the left. The more stable GFP marker is weakly detected in Pdm1+ nuclei, presumably the ICPs (some indicated by arrows in A’-A’’’) in the center of the midgut sac. Arrowheads in A” indicate locations of Hey-GFP+ Pdm1- cells representing EEs. (C-C’’’) Horizontal view of a stage 16 embryo. Anterior is at the top. ECs of anterior most domain of the midgut primordium are stained for Hey-GFP expression (some indicated by arrows in C’-C’’’). Arrowheads in C’’ point to positions of HeyGFP+ Pdm1- cells representing EEs in two of the midgut primordium domains. Note the strong Hey-GFP expression at the brain lobes at the anterior (up) of the embryo in C. Scale bar, 40μm (A-C), 34μm (B’-B”’), 20μm (Α’-A”’, C’-C”’).
Hey is a target of Notch signaling in the embryonic midgut primordium
Active Notch signaling is required for proper midgut development and cell fate decisions between enterocytes and enteroendocrine/AMP precursor populations (Tepass and Hartenstein 1995; Takashima et al. 2011). Even as early as stages 10-11 when the mesenchymal cells of amg and pmg rudiments undergo proliferation and migration, Notch activation favors EC fate in cells that are in contact with the underlying visceral musculature and go through MET. Thus, in wild type embryos with balanced Notch activation, the majority of the endodermal cells become ECs and only a small percentage of them residing in the inner layer of the migrating cell populations will remain undifferentiated (AMP) or will follow the EE fate. By contrast, in embryos with mutations in either Notch or Delta (Dl) signaling molecules, all the cells express the EE marker Prospero, resulting in a massive increase of AMP/EE progenitors at the expense of the EC population (Tepass and Hartenstein 1995; Takashima et al. 2011). Whether Notch signaling is involved in any additional cell fate choice(s) that perhaps take place in later embryonic stages to generate distinct post mitotic cells of the midgut epithelium has yet to be documented.
Hey is a target of Notch in many CNS nascent neurons (Monastirioti et al., 2010), so we wanted to address whether this is also the case in the midgut. Based on our data, Hey is expressed in the embryonic midgut primordium in time periods later than those of stage 10-12, where intense Notch signaling has been documented, as mentioned above. Hey expression identified almost half of the postmitotic Pros+ cells that switch off escargot at stage 14/15 and become EE cells, thus we questioned whether this expression is regulated by a later pulse of Notch signaling or it is Notch independent, as has been suggested for Hey expression in the adult midgut (Flint Brodsly et al., 2019). In order to address this issue, we monitored Hey immunostaining in embryos bearing mutations with severe defects on activation of the Notch pathway.
In double-null mutant embryos for Dl and Ser, the genes encoding the only two Notch ligands in Drosophila, establishment of EC fate is disrupted in similar fashion to that observed for Dl mutants alone (Takashima et al., 2011). All endodermal cells express Prospero, and the midgut primordium develops with severe malformation (Fig. 6D). Despite displaying a large increase in total numbers of Pros-positive cells (Fig. 6D), the midguts of Dl Ser embryos are completely devoid of Hey expression (Fig. 6 D-D’), suggesting that Hey is indeed a Notch target in the developing midgut. We also examined Df(3R)E(spl)b32.2 embryos bearing a deficiency for the entire E(spl) complex. This includes seven bHLH-O genes that are expressed in response to Notch, are mediators of Notch activity in many instances, but are not required for Notch signal transduction per se. Similarly to Dl Ser embryos, Df(3R)E(spl)b32.2 homozygous embryos display a malformed midgut with an increased Pros+ population (Fig. 6 E-E’); in this case, however, parallel to the increased number of Pros-positive cells, the number of Hey-positive cells is also increased (Fig. 6 E-E’’). This observation supports the notion that Hey expression in the EE subpopulation of the developing midgut is responsive to Notch signaling but is independent of E(spl), similarly to what we have previously described for its expression in the embryonic CNS (Monastirioti et al., 2010). We compared Hey expression with that of another common Notch target, E(spl)m8, using a genomic duplication of E(spl)m8 tagged with GFP (Kudron et al., 2018). Whereas E(spl)-m8GFP appears in amg/pmg primordia as early as stages 10-12 (Fig. S2 A,B) before any Hey expression, they are highly co-expressed at stages 13/14 (Fig. S2 C-C’’). E(spl)m8 disappears from the Hey-positive EE subpopulation by stage 15/16, although Hey remains (Fig S2B).
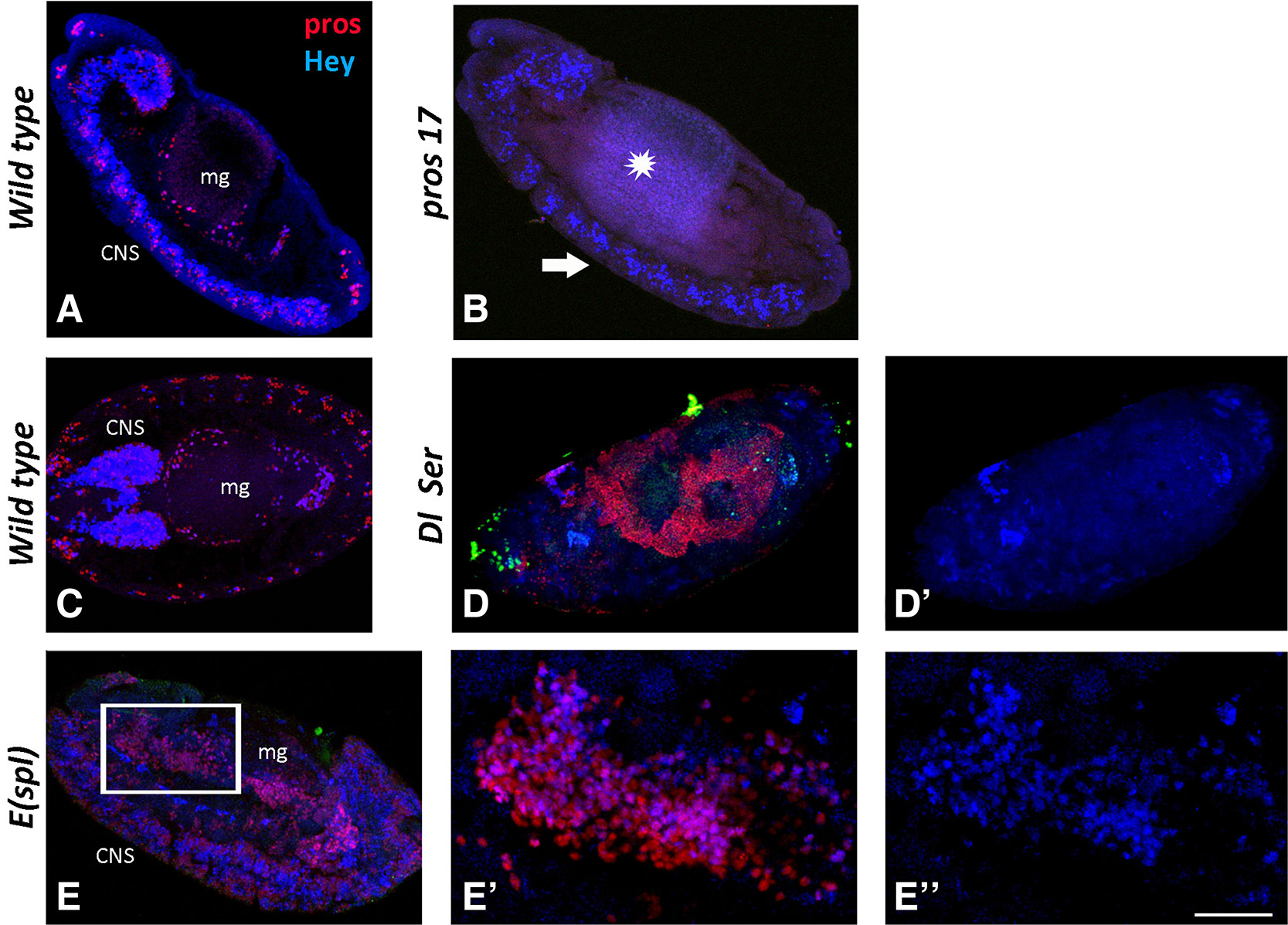
Fig. 6. Hey expression in midgut primordium of mutant backgrounds.
Wild type and mutant embryos stained for Prospero (red) and Hey (blue). (A, C) Sagittal view of a stage 13 (A) and horizontal view of a stage 15 (C) wild type embryo. Prospero (red) and Hey (blue) expression is detected in CNS and midgut primordium (mg). Anterior is at the top in (A) and to the left in (C). (B, D, E) Sagittal view of embryos from different mutant backgrounds. (B) In pros17 mutants, Prospero immunoreactivity (red) is not detected in either the midgut primordium (star) or in the CNS (arrow). Hey expression (blue) is disturbed in the CNS (note that fewer Hey-positive cells are detected), while it is completely missing from the midgut. In Dl Ser mutant embryo (D-D’), there is a massive excess of Pros+ (red in D) cells that also displays the malformation of the midgut primordium. Hey expression (blue, D’) is absent from both tissues, indicating that it is Notch dependent. (E-E’’) Transduction of Notch signal is normal in embryos of E(spl) mutant background, but some of its downstream effects are abolished due to the loss of the E(spl) effectors. Thus, E(spl) mutants display a CNS hyperplasia (E) and a malformed midgut with excess of Pros+ (red) cells (E, E’) similarly to Dl Ser mutants. Hey expression (blue) in this mutant background is not affected, and it is evident in many more neurons in the hyperplastic CNS (E) as well as in many more Pros+ cells in the midgut (E’, E’’) compared with wild type. (E) and (E’’) are higher magnification images of the boxed area in (E). Scale bar, 40μm (A-E, D’), 17μm (E’-E’’).
We also examined Hey expression in embryos mutant for prospero. We found that pros17 null embryos lacked Hey immunoreactivity, suggesting that expression of Prospero is a prerequisite for expression of Hey in the EE cells (Fig. 6B)
Hey expression in Drosophila larval midgut
Morphogenesis of Drosophila larva midgut is complete by the end of embryogenesis, so that hatching larvae are provided with a functional midgut required for digestion of the consumed food. The three different cell populations of the larva midgut epithelium - ECs, EEs, and AMPs - that have been already established by late embryonic stages (st16-17) can be distinguished by both their specific markers Pdm1, Prospero and Escargot, respectively, and by the size of their nucleus, large for polyploid ECs versus small for diploid EEs and AMPs. In particular, the EE cell population appears as single cells scattered along midgut epithelium throughout larva life, while AMPs are initially single scattered cells during 1st instar larva, while later on they undergo several rounds of mitosis forming characteristic groups of 2-3 cells (2nd instar) or 7-8 cells (late 3rd instar), called AMP islands ((Jiang and Edgar, 2009).
We initially investigated the expression of Hey in the larval midgut by immunostaining with the same specific antibody against Drosophila Hey (Monastirioti et al., 2010) used in our embryo experiments. In contrast to the robust expression of Hey in a large subpopulation of Pros+ EE cells during embryonic development, only poor Hey expression was observed in the larval midgut. Only in very early 1st larval (L1) stage, a couple of hours after hatching, did we manage to detect weak Hey immunostaining in very few Pros+ EE cells as well as in a few ECs in the very anterior region next to proventriculus (not shown). These observations are consistent with our hypothesis that robust Hey expression takes place in early EE specification, while later on, in the mature larval midgut, Hey is either not expressed or its expression is maintained at such low levels that are beyond detection capacity of our antibody.
In an attempt to clarify this issue, we took advantage of a recently generated transgenic line that expresses a GFP-tagged Hey protein from a large genomic transgene (Kudron et al., 2018). This fusion lies in the Hey locus, therefore its expression is regulated as the endogenous gene, and GFP follows the characteristic expression pattern of Hey. Given the stability of GFP protein, it was considered a good tool for following putative low Hey expression during larval stages. We first examined the expression of the Hey-GFP fusion protein in the embryonic midgut by specific antibodies against both Hey and GFP. During embryonic stages, all cells with robust Hey expression are detected by both a-Hey and a-GFP antibodies (Fig. S3). Moreover, we were able to detect GFP expression in the group of the presumptive ICPs identified by the a-Hey specific antibody in stages 11-14 (Fig. 5 A-A’’’, Fig. S3); to our surprise, weak but clear GFP immunostaining was also evident in few ECs in the most anterior midgut sac of stage 16 embryos (Fig. 5 C-C’’), where a-Hey antibody cannot detect the endogenous protein. The results confirmed the validity of the Hey-GFP tool for monitoring Hey expression, and indicated its potential to detect putative low expression through GFP stability.
Subsequently, we examined the expression of the Hey-GFP fusion in early L1 and 3rd instar larval (L3) stages by antibodies against GFP (Fig. 7). In anterior regions of the L1 midgut, Hey-GFP+ cells were detected among the Pros+ EE population as well as in the majority of ECs (Fig. 7, midgut lumen and surface panels). Weak Hey-GFP expression, though, was also evident in a few Pros+ EE cells in L3 stage (not shown). However, when we treated 3rd instar midguts with methanol during fixation (which helps to strengthen low signals by better penetration of the tissue), strong GFP immunostaining became evident in considerably more EE cells throughout the midgut, and weaker but quite evident GFP was observed in ECs of anterior/middle regions (Fig. 8 A-A’’), as in younger larvae. The above manipulations support the notion that Hey expression levels in larvae midguts are too low to be detected by the a-Hey antibody.
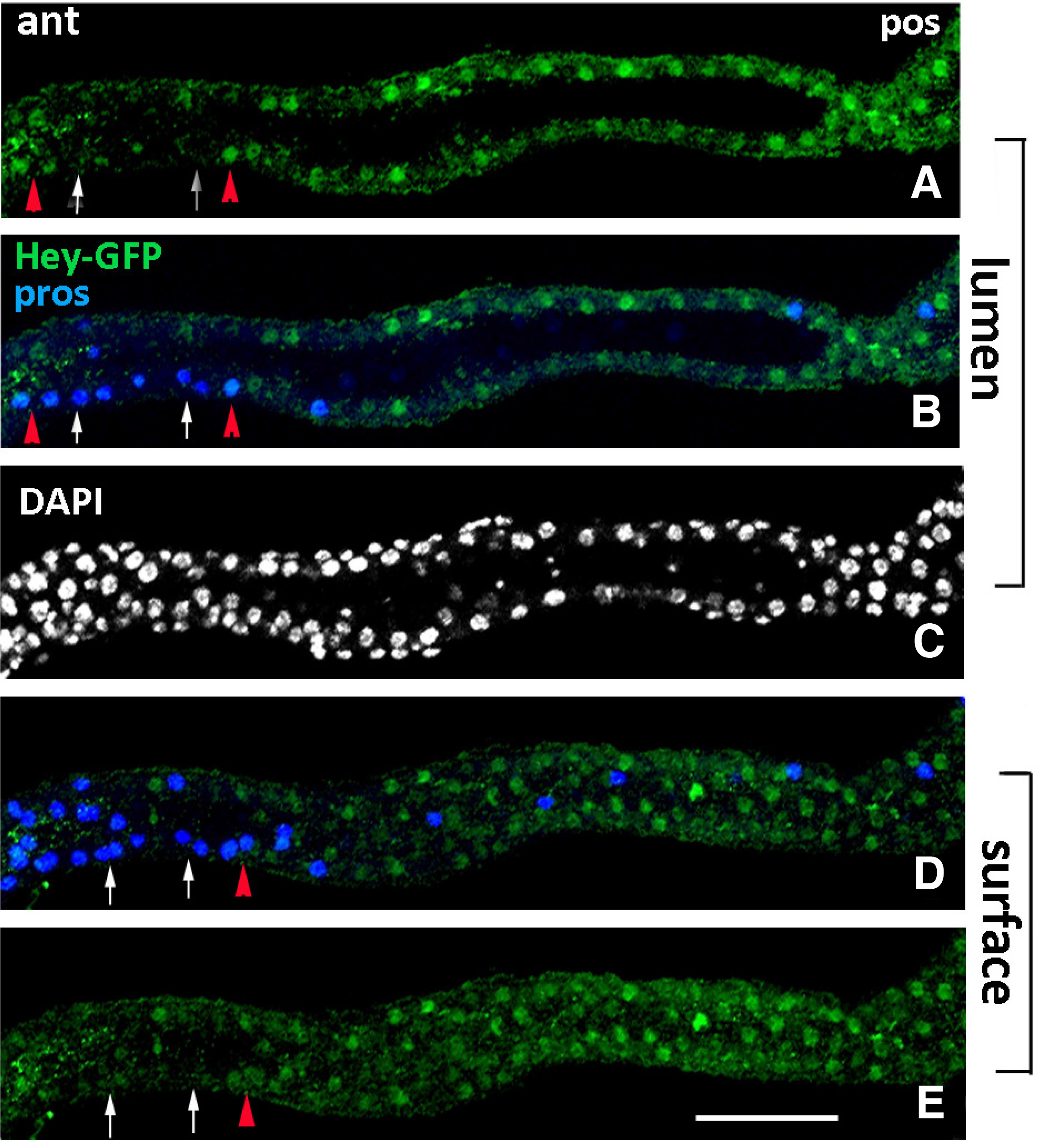
Fig. 7. Hey expression in early larval midgut.
Early 1st instar larval gut of Hey-GFP line stained for GFP (green), indicating expression of the Hey-GFP fusion and for Prospero (blue). All images represent group of sections from the same anterior midgut region focused on different levels of the tissue, lumen (A-C) and surface (D-E). Hey-GFP expression (green) is detected in a subset of Pros+ (blue) cells in this early stage (examples shown by red triangles in A, B, D, E). White arrows in the same images point to Pros+ cells that are devoid of Hey-GFP expression. Many enterocytes are also stained for Hey-GFP (green) along the epithelium (A-E). Anterior is left. Scale bar, 25μm.
In addition to the Pros+ single nuclei, Hey-GFP fusion expression was often observed in a single cell of a subset of AMP islands, primarily at the posterior midgut (Fig. 8 B-C) of late 3rd instar larvae. Hey-GFP staining was detected only in AMP islands that included Pros+ cell doublets (Fig. 8 B’’,C’’), and in fact in one of these two cells (Fig. 8 B’’-B’’’,C’’-C’’’). We also observed the same with Hey immunostainings, which indicates that Hey expression in this cell is strong enough to be detected with the a-Hey specific antibody. Such de novo expression of Prospero within late L3 AMP islands has also been observed before (Takashima et al., 2011). It is conceivable that these Pros+ Esg+ AMPs represent EE precursors that have undergone an asymmetric division, as marked by Hey expression in one of them, and that they generate the presumptive pupal EE cells suggested previously (Takashima et al., 2011).
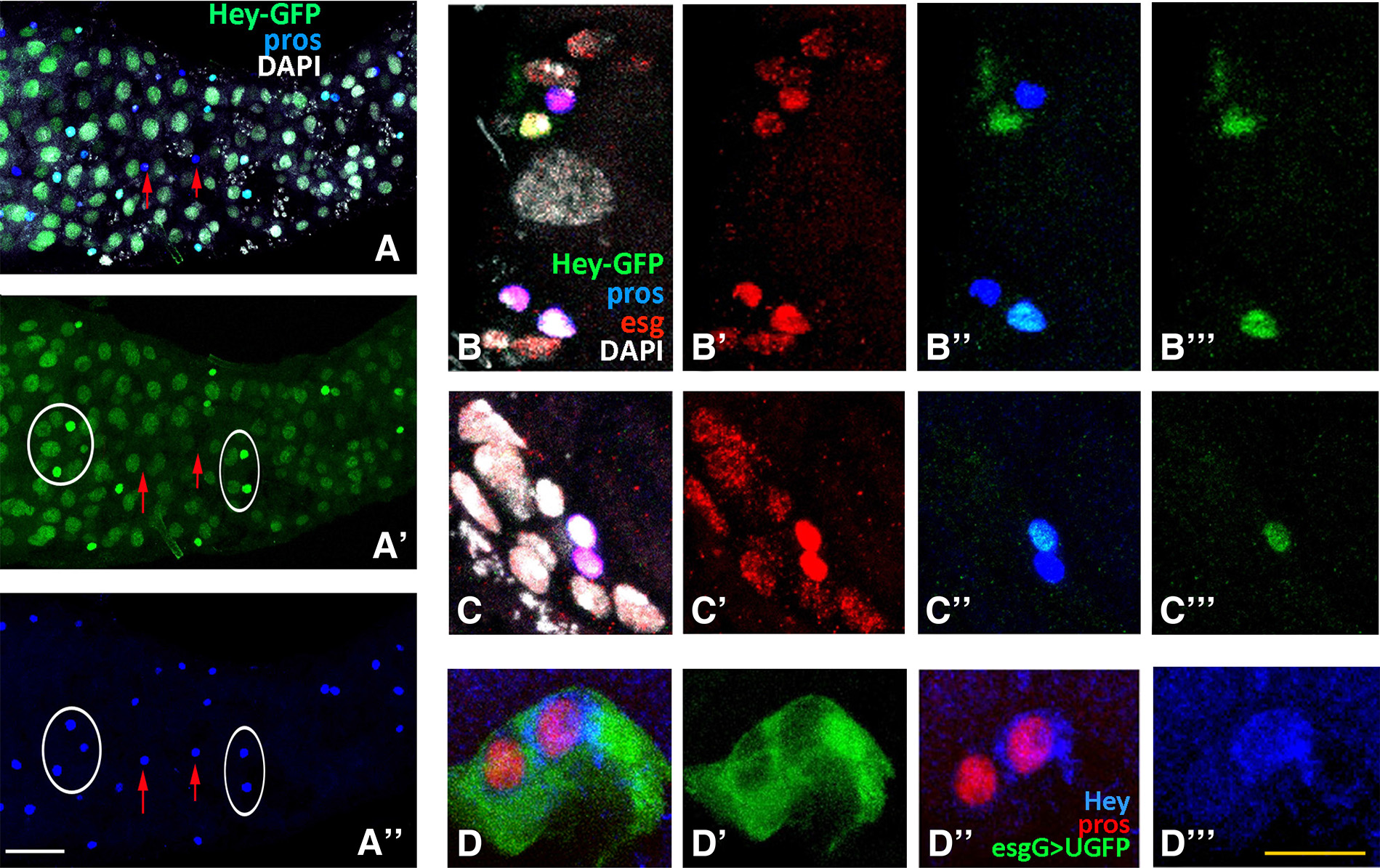
Fig. 8. Hey expression in late larval midgut.
Third instar larval gut of Hey-GFP line (A-C), treated with methanol fixation and stained for GFP (green), indicating expression of the Hey-GFP fusion, for Prospero (blue) and Escargot (red). (A-A’’) Images of sections from the same anterior midgut surface region. Many enterocytes are stained for Hey-GFP (green) along the epithelium (A, A’). Hey-GFP expression (green) is also detected in a subset of Pros+ (blue) cells (examples shown by ovals in A’, A’’). Red arrows in the same images point to Pros+ cells that are devoid of Hey-GFP expression. Anterior is to the left. (B-C) Representative images of posterior midgut AMP islands showing the de novo Prospero expression (blue) in two AMP cells (B’’, C’’). One of the two Pros+ cells is also stained for Hey-GFP (green) (B’’’, C’’’). DAPI nuclear staining (white) and Escargot immunoreactivity (red) are detected in all AMP cells of the island. Note that the Pros+ Hey-GFP+ doublet is strongly stained by Escargot (B’, C’). (D-D’’’) Example of an AMP island at the posterior region of third instar lava midgut from the esgGAL4>UGFP line stained with antibodies against GFP (green), displaying the AMP island cells, Prospero (red) and Hey (blue). One cell of the Pros+ pair (red, D’’) within the island (D’) is Hey-positive as well (D’’’), suggesting adequate levels of Hey protein expressed to be detected by Hey immunostaining. Scale bars, 45μm (A-A”), 12μm (B-C”’), 10μm (D-D”’).
In conclusion, the above data indicate that throughout larval life, Hey is still expressed in a subpopulation of EE cells, in particular a subpopulation born within AMP islands during late third instar. Hey is also expressed in some ECs of the anterior midgut, albeit in very low levels.
Discussion
In this study, we investigated the expression pattern of Drosophila Hey in a non-neuronal tissue during embryogenesis. We found that Hey is expressed in cells of the midgut primordium that originates from the invaginated endoderm in the germ band extended embryo and develops into the functional midgut epithelia of the hatching larvae. Three distinct types of midgut cells are established through embryogenesis (ECs, EEs and AMPs), and our study shows that Hey expression primarily marks cells of the EE fate, in fact approximately half of the latter. Having examined a large number of embryos spanning all stages of embryogenesis, we did not detect any additional site of Hey expression besides the nervous system and midgut, although we cannot exclude the possibility of expression below the level of detection of our antibody.
At the onset of its expression (stage 11-12), Hey is weakly and transiently expressed in large nuclei in the posterior midgut rudiment, which we hypothesize to be a subset of the interstitial cell precursors (ICPs). From stage 13 onwards, Hey is strongly detected in a subpopulation of small nuclei displaying immunostaining for Prospero, suggesting that they possess enteroendocrine character. Hey immunoreactivity never coincides with phospho-H3, suggesting that it is expressed in postmitotic EE cells. This is in contrast with Prospero, which is occasionally seen expressed in cells undergoing mitosis. Taking into account the fact that Hey marks about half of the Pros-positive EE cells at stage 15 (when mitotic activity in the midgut has ceased), we propose that Hey is switched on in one of the two sibling EE cells that arise after the asymmetric division of a Pros-positive / Hey-negative EE precursor. This asymmetric division takes place around stage 12-13 of embryogenesis at a time when the entire midgut is still positive for Esg-GFP, which marks undifferentiated cells; however, soon afterwards, by stage 15, Esg-GFP expression is extinguished in most cells, including all Pros-positive EEs, and remains only in the adult midgut precursors (AMPs). The proposed EE precursor division must rely on asymmetric Notch signaling to distinguish between the two sibling EE fates, since Hey expression in the midgut is lost in Notch pathway mutants. This is analogous to the Notch dependence of Hey expression in the developing CNS, where it is turned on in one of the two sibling neurons born after an asymmetric GMC division. EE precursor asymmetric divisions have also been reported to take place in the pupal and adult midgut (Beehler-Evans and Micchelli 2015; Guo and Ohlstein 2015), but we have not examined these stages for Hey expression in the present study.
A presumptive asymmetric division may also take place in late larva midgut. Takashima et al. (2011) had observed a new set of EE cells expressing Prospero that, contrary to the scattered larva EE cells, are smaller, reside within AMP islands and during pupariation increase in number; they integrate into the transient pupal midgut and they are lost along with the latter upon eclosion. We also observed such cells within a small number of AMP islands in late larva midguts as doublets of Pros+ cells, one of which displayed Hey immunoreactivity. Although we have not directly tested for Notch dependence, the presence of Hey-positive/negative Pros-positive doublets is consistent with a Notch/Hey asymmetric cell division underlying the de novo formation of EE cells that takes place at the larva to pupa transition phase.
Besides the robust Hey expression in EE subpopulation of the developing larval midgut, weak expression was noticed in broad regions of the newly hatched 1st instar larva midgut. This was presaged by Hey expression in scattered large nuclei corresponding to Pdm1+ enterocytes at very late embryonic stages (stage 16). Their location was in the anterior most of the four midgut primordium domains, which gives rise to the proventriculus, gastric cecae and anterior most region of larval midgut (Lemaitre and Miguel-Aliaga, 2013). Accordingly, weak Hey expression was detected in anterior regions - between proventriculus and copper cell region of the midgut (unpublished observations) - during all larval stages. We have not investigated this expression further, but we hypothesize that it may be analogous to the Notch independent expression of Hey described before (Flint Brodsly et al., 2019), required for integrity and maintenance of the adult midgut enterocytes. The use of a Hey-GFP large genomic transgene (Kudron et al., 2018) increases the sensitivity of detection of such weakly expressing cells, presumably via the stability of the Hey fusion protein.
Our study reveals that the Drosophila Hey bHLH-O transcription factor, a target and effector of activated Notch pathway, exhibits a Notch dependent novel expression pattern during development of the larval midgut in a subset of newly born postmitotic EE cells, and this pattern resembles its expression mode in newly born neurons (Monastirioti et al., 2010). It is of interest to this study that earlier reports have suggested that generation of enteroendocrine cells of the adult Drosophila is analogous to the generation of neuronal lineages (Guo and Ohlstein 2015; Hartenstein et al. 2017), as it involves the expression of proneural genes (Bardin et al., 2010) and asymmetric divisions regulated by Notch signaling. By interfering with a single transcriptional repressor, Tramtrack, it is even possible to convert intestinal lineages into neural-like cells, as has recently been demonstrated (Li et al., 2020). This implies that chromatin-based switches may be superimposed on the same regulatory cassette, like the Notch/Hey module, to produce different developmental outcomes in different contexts.
Materials and Methods
Fly strains
All fly stocks used for the immunohistochemical experiments were obtained from the Bloomington Stock Centre, VDRC (Vienna Drosophila Resource Center), or individual laboratories. The Hey-GFP (Kudron et al., 2018) and E(spl)m8-GFP (Piwko et al., 2019) lines were used to detect Hey-positive and E(spl)m8-positive cells, respectively. The esg-Gal4; UAS-mCD8GFP (Micchelli and Perrimon, 2006) and esg-GFP/ TM3 sb (Sarov et al., 2016) lines were used to identify AMP islands in 3rd instar larva midguts and AMP cells of the embryonic midgut primordia, respectively.
Mutant backgrounds were: e Df(3R)E(spl)b32.2/TM3, w; FRT82B Dlrev10 e SerRX106/ TM3 twistG> UGFP TM6B, w; pros17/twistG>UGFP.
Larval Staging
Larvae of specified stage were obtained from 2-3 hours embryo collections in 25 °C. The embryos were aged appropriately to obtain late L1 (48h AED), and “wandering” late L3 (108-119h AED) instar larvae [AED: After Egg Deposition].
Immunohistochemistry
Fixation and immunohistochemistry of embryos was performed according to standard protocols. For larval midgut immunohistochemistry, only female larva midguts were dissected, fixed in 4% formaldehyde (FA) in 1xPBS for 20 min at room temperature (RT) and quickly washed in 1xPBS. When methanol treatment was included after FA fixation, 100% methanol was added dropwise till PBS: Methanol ratio was 1:2 and then the mixture was substituted by methanol 100% for an incubation of 5min in RT. Subsequently, PBS was added dropwise until a PBS: Methanol ratio 2:1 and tissues were washed in PBS (x3) followed by a quick wash in PT (1xPBS, 0.2% Triton). For blocking and permeabilization, midguts were treated with PBT (PT+ 0.5% BSA) for at least 30 min at room temperature. Incubation in primary antibodies diluted in PBT was conducted O/N (4 °C), followed by washes in PBT and incubation in secondary antibodies for 2 hours at room temperature. After extensive washes in PT and a quick wash in PBS, DAPI incubation was performed for five minutes to stain all cell nuclei. Finally, tissues were washed in PBS and mounted in n-propyl gallate / glycerol. Embryos and midgut tissues were imaged in TCS SP8 confocal microscope system (Leica) and image analysis was performed with the Leica LAS X software.
Primary antibodies used were: Guinea pig a-Hey, 1:1000 (Monastirioti et al., 2010); mouse a-Prospero MR1A, 1:20 (DSHB); rabbit a-GFP, 1:30,000 (Minotech); mouse a- Nubbin (Pdm1), 1:5 (gift from M. Averof), rat a-escargot, 1:1000; rabbit a-PH3, 1:2000 (Milipore). Detection was done using secondary antibodies conjugated to Alexa 488, 555, 568, 633 or 647 (Molecular Probes), or Cy3, Cy5 (Jackson ImmunoResearch).
Supplementary Material
Acknowledgements
We thank Michalis Averof for his kind gift of Nubbin (Pdm1) antibody and Chrysoula Pitsouli for the esg-Gal4; UAS-mCD8GFP strain. We also thank Margarita Stapountzis for confocal training to E. Skafida and for support and technical assistance in the confocal unit. The work was funded by a Fondation Santé grant (C.D.) and intramural Institute Biology and Biotechnology (IMBB) support (M.M.).
Abbreviations
amg, anterior midgut ; AMPs, adult midgut progenitors ; bHLH, basic-helix-loop-helix ; CNS, central nervous system ; GMC, ganglion mother cell ; EC, enterocyte ; EE, enteroendocrine ; EMT, epithelial-to-mesenchymal transition ; ICPs, interstitial cell precursors ; MET, mesenchymal-to-epithelial transition ; PMECs, principal midgut epithelial cells ; pmg, posterior midgut ;References
Bardin A. J., Perdigoto C. N., Southall T. D., Brand A. H., Schweisguth F. (2010). Transcriptional control of stem cell maintenance in the Drosophila intestine . Development 137: 705-714.
Beehler-Evans R., Micchelli C. A. (2015). Generation of enteroendocrine cell diversity in midgut stem cell lineages. Development 142: 654-664.
Campbell K., Casanova J. (2015). A role for E-cadherin in ensuring cohesive migration of a heterogeneous population of non-epithelial cells. Nature Communications 6: 7998.
Choksi S. P., Southall T. D., Bossing T., Edoff K., de Wit E., Fischer B. E., van Steensel B., Micklem G., Brand A. H. (2006). Prospero Acts as a Binary Switch between Self-Renewal and Differentiation in Drosophila Neural Stem Cells. Developmental Cell 11: 775-789.
Dubreuil R.R., Frankel J., Wang P., Howrylak J., Kappil M., Grushko T.A. (1998). Mutations of α Spectrin andlabialBlock Cuprophilic Cell Differentiation and Acid Secretion in the Middle Midgut ofDrosophilaLarvae. Developmental Biology 194: 1-11.
Dutta D., Dobson A. J., Houtz P. L., Gläßer C., Revah J., Korzelius J., Patel P. H., Edgar B. A., Buchon N. (2015). Regional Cell-Specific Transcriptome Mapping Reveals Regulatory Complexity in the Adult Drosophila Midgut. Cell Reports 12: 346-358.
Fischer A., Gessler M. (2007). Delta Notch and then? Protein interactions and proposed modes of repression by Hes and Hey bHLH factors. Nucleic Acids Research 35: 4583-4596.
Flint Brodsly N., Bitman-Lotan E., Boico O., Shafat A., Monastirioti M., Gessler M., Delidakis C., Rincon-Arano H., Orian A. (2019). The transcription factor Hey and nuclear lamins specify and maintain cell identity. eLife 8: e44745.
Guo Z., Ohlstein B. (2015). Bidirectional Notch signaling regulates Drosophila intestinal stem cell multipotency . Science 350: 6263.
Hartenstein A.Y., Rugendorff A., Tepass U., Hartenstein V. (1992). The function of the neurogenic genes during epithelial development in the Drosophila embryo. Development 116: 1203-1220.
Hartenstein V., Takashima S., Hartenstein P., Asanad S., Asanad K. (2017). bHLH proneural genes as cell fate determinants of entero-endocrine cells, an evolutionarily conserved lineage sharing a common root with sensory neurons. Developmental Biology 431: 36-47.
Hartenstein V., Wodarz A. (2013). Initial neurogenesis in Drosophila . Wiley Interdisciplinary Reviews: Developmental Biology 2: 823-823.
Heisig J., Weber D., Englberger E., Winkler A., Kneitz S., Sung W.K., Wolf E., Eilers M., Wei C.L., Gessler M. (2012). Target Gene Analysis by Microarrays and Chromatin Immunoprecipitation Identifies HEY Proteins as Highly Redundant bHLH Repressors. PLoS Genetics 8: e1002728.
Jiang H., Edgar B. A. (2009). EGFR signaling regulates the proliferation of Drosophila adult midgut progenitors . Development 136: 483-493.
Kudron M. M., Victorsen A., Gevirtzman L., Hillier L.D. W., Fisher W. W., Vafeados D., Kirkey M., Hammonds A. S., Gersch J., Ammouri H., Wall M. L., Moran J., Steffen D., Szynkarek M., Seabrook-Sturgis S., Jameel N., Kadaba M., Patton J., Terrell R., Corson M., Durham T. J., Park S., Samanta S., Han M., Xu J., Yan K.K., Celniker S. E., White K. P., Ma L., Gerstein M., Reinke V., Waterston R. H. (2018). The ModERN Resource: Genome-Wide Binding Profiles for Hundreds of Drosophila and Caenorhabditis elegans Transcription Factors . Genetics 208: 937-949.
Lee T. (2017). Wiring the Drosophila Brain with Individually Tailored Neural Lineages. Current Biology 27: R77-R82.
Lemaitre B., Miguel-Aliaga I. (2013). The Digestive Tract of Drosophila melanogaster . Annual Review of Genetics 47: 377-404.
Li Z., Guo X., Huang H., Wang C., Yang F., Zhang Y., Wang J., Han L., Jin Z., Cai T., Xi R. (2020). A Switch in Tissue Stem Cell Identity Causes Neuroendocrine Tumors in Drosophila Gut. Cell Reports 33: 108459.
Lu H., Kozhina E., Mahadevaraju S., Yang D., Avila F. W., Erickson J. W. (2008). Maternal Groucho and bHLH repressors amplify the dose-sensitive X chromosome signal in Drosophila sex determination. Developmental Biology 323: 248-260.
Micchelli C. A., Perrimon N. (2006). Evidence that stem cells reside in the adult Drosophila midgut epithelium. Nature 439: 475-479.
Monastirioti M., Giagtzoglou N., Koumbanakis K. A., Zacharioudaki E., Deligiannaki M., Wech I., Almeida M., Preiss A., Bray S., Delidakis C. (2010). Drosophila Hey is a target of Notch in asymmetric divisions during embryonic and larval neurogenesis . Development 137: 191-201.
Nakagoshi H. (2005). Functional specification in the Drosophila endoderm. Development, Growth and Differentiation 47: 383-392.
Overend G., Luo Y., Henderson L., Douglas A. E., Davies S. A., Dow J. A. T. (2016). Molecular mechanism and functional significance of acid generation in the Drosophila midgut. Scientific Reports 6: 27242.
Pitsidianaki I., Morgan J., Adams J., Campbell K. (2021). Mesenchymal-to-epithelial transitions require tissue-specific interactions with distinct laminins. Journal of Cell Biology 220: e202010154.
Piwko P., Vitsaki I., Livadaras I., Delidakis C. (2019). The Role of Insulators in Transgene Transvection in Drosophila . Genetics 212: 489-508.
Sarov M., Barz C., Jambor H., Hein M. Y., Schmied C., Suchold D., Stender B., Janosch S., KJ V. V., Krishnan R.T., Krishnamoorthy A., Ferreira I. R.S., Ejsmont R. K., Finkl K., Hasse S., Kämpfer P., Plewka N., Vinis E., Schloissnig S., Knust E., Hartenstein V., Mann M., Ramaswami M., VijayRaghavan K., Tomancak P., Schnorrer F. (2016). A genome-wide resource for the analysis of protein localisation in Drosophila. eLife 5: e12068.
Skeath J.B., Doe C.Q. (1998). Sanpodo and Notch act in opposition to Numb to distinguish sibling neuron fates in the Drosophila CNS. Development 125: 1857-1865.
Stampfel G., Kazmar T., Frank O., Wienerroither S., Reiter F., Stark A. (2015). Transcriptional regulators form diverse groups with context-dependent regulatory functions. Nature 528: 147-151.
Takashima S., Adams K. L., Ortiz P. A., Ying C. T., Moridzadeh R., Younossi-Hartenstein A., Hartenstein V. (2011). Development of the Drosophila entero-endocrine lineage and its specification by the Notch signaling pathway. Developmental Biology 353: 161-172.
Takashima S., Hartenstein V. (2012). Genetic Control of Intestinal Stem Cell Specification and Development: A Comparative View. Stem Cell Reviews and Reports 8: 597-608.
Tepass U., Hartenstein V. (1994). Epithelium formation in the Drosophila midgut depends on the interaction of endoderm and mesoderm. Development 120: 579-590.
Tepass U., Hartenstein V. (1995). Neurogenic and proneural genes control cell fate specification in the Drosophila endoderm. Development 121: 393-405.
Weber D., Wiese C., Gessler M. (2014). Hey bHLH Transcription Factors. In bHLH Transcription Factors in Development and Disease. (Ed. Taneja, R., ) Academic Press.