Int. J. Dev. Biol. 66: 177 - 186 (2022)
Special Issue: Developmental Biology in Greece
RNAi silencing of the Arabidopsis thaliana ULCS1 gene results in pleiotropic phenotypes during plant growth and development
Original Article | Published: 9 August 2021
Abstract
WD40-repeat-containing proteins (WDRs) are highly abundant in all eukaryotes. Several have been implicated as subunits of multi-protein CRL E3 ligase complexes that regulate ubiquitination mediated protein degradation and thus various cellular and developmental processes. Impairment of the WDR protein ULCS1 from Arabidopsis causes pleiotropic phenotypes during plant development, including reduced lignification, anther indehiscence, and sterility. Here we show that RNAi-mediated downregulation of ULCS1 results in a fast-growing phenotype during vegetative development. Due to accelerated growth, ulcs1i mutants reach their vegetative to reproductive transition point earlier than WT plants. However, their comparable germination rate and their similar number of secondary branches and rosette leaves at bolting indicate that ulcs1i is not an early flowering time mutant. GUS staining of progeny, obtained from crosses between ulcs1i and CYCB1::GUS plants, revealed an increased number of mitotic cell divisions in the root meristems of ulcs1i compared to WT. Immunolabeling of homogalacturonans (HGAs) epitopes showed significant fluorescent signal differences at the cell walls and the mucilage of the seeds between ulcs1i and WT. Furthermore, we demonstrate that ULCS1 interacts with the UBA-like protein in a yeast two-hybrid assay, suggesting a direct or indirect physical coupling of these proteins in Arabidopsis.
Keywords
arabidopsis, CRL complex, ubiquitination, WD40 motif, WDR protein, vegetative growth, seed coat, mucilage, homogalacturonans
Introduction
WDR proteins (WD40 protein domain, or beta-transducin repeat-WDRs) constitute a large prokaryotic and eukaryotic family and act as scaffold subunits for the assembly of multi-protein complexes (Neer et al., 1994, van Nocker et al., 2003; Jain and S. Pandey 2018). They are characterized by the presence of a repetitive, short structural motif of about 44-60 amino acids, which contains a conserved glycine-histidine (GH) dipeptide in the N-terminus and a signature tryptophan-aspartic (WD) dipeptide in the C-terminus of the domain (Fong et al., 1986; Smith et al., 1999). Most WDR proteins contain 4 to 16 tandem WD40 repeats (WD40 domain) and fold into a tertiary β-propeller architecture that can provide a platform for protein complex assembly (Lambright et al., 1996). The WD40 domain was initially identified in the Gβ subunit of the heterotrimeric G receptor-signaling pathway, but is now considered to be one of the most abundant domains in eukaryotic proteins. Although more than 4,000 WD40 domain-containing proteins have also been identified in bacteria, primarily Cyanobacteria and Planctomycetes, they are much less abundant in most prokaryotes (Hu et al., 2017). Since no WD40 domain has yet been identified to possess an intrinsic enzymatic activity, evidence suggests that WDRs act as scaffold proteins for the assembly of large molecular machines, such as ubiquitin ligase and histone methylation complexes. They play central roles and can regulate various environmental stress responses or developmental processes, by acting as hubs for protein-protein or protein-nucleic acid interactions (Stirnimann et al., 2010; Jain and S. Pandey 2018). It is generally accepted that WDR proteins can dock a variety of substrates, with similar or distinct modes, by utilizing the entire surface of their β-propeller architecture. The enormous variety of WDR proteins and their sequence diversity outside of the WD40 repeats, which determines the specificity of the protein, enables them to coordinate the assembly of various multi-protein complexes. Thus, WDRs act as key regulators in numerous cellular processes and networks, such as signal transduction, mRNA synthesis, histone methylation, protein ubiquitination, cell cycle control, cytoskeletal organization, apoptosis, and development (Yu et al., 2000; Xu and Min, 2011; Zhang and Zhang, 2015; Jain and S. Pandey 2018).
A recent genome-wide analysis revealed that the Arabidopsis thaliana proteome contains ca. 269 low and high molecular weight WDR proteins, which can be categorized into 143 distinct families (Li et al., 2014). All of these contain at least one WD40 repeat, while several members have previously been shown to be subunits of regulatory complexes involved in diverse developmental processes, such as meristem organization (Kaya et al., 2001), embryo development (Bjerkan et al., 2012), differentiation of epidermal cells (Wang et al., 2016) or pollen (Xu et al., 2011), as well as adaptation to long day lengths and flowering time control (Inouchi et al., 2000; Jiang et al., 2011; Kapolas et al., 2016). Several of these proteins are members of the ubiquitin-proteasome system (UPS). The UPS leads to ubiquitination of target proteins, which thereafter are transported to the vacuole for degradation by the 26S proteasome (Scheuring et al., 2012). This process is mediated by the sequential action of the E1 ubiquitin-activating enzymes, the E2 ubiquitin-conjugating enzymes, to which the ubiquitin is transferred from the E1, and the E3 ubiquitin ligase complexes (E3s) that catalyze the transfer of the ubiquitin to the target protein (Nelson and Millar, 2015). Amongst the identified E3s are the cullin-RING ligases (CRLs), three of which are recognized in plants: the SCF, BTB, and DWD E3s (Hua and Vierstra, 2011; Chen and Hellmann, 2013). Besides being implicated in, for example, embryo development (Figueroa et al., 2005), light signaling (Chen et al., 2006), pollen development (Gusti et al., 2009), and secondary growth (Beris et al., 2016), impaired CRL function has also been related to pleiotropic phenotypes and other developmental defects (Lechner et al., 2002; Chantha et al., 2007).
We have previously shown that the ULCS1 gene from A. thaliana encodes for a small molecular weight WDR protein and interacts with the E3 Cullin Ring Ligase subunit DDB1a, probably regulating the degradation of proteins involved in secondary wall modifications (Beris et al., 2016). RNAi-mediated silencing mutants (ulcs1) exhibit a reduced deposition of secondary cell wall components and lignin, both in the vascular system of stems and in anthers, resulting in anther indehiscence and sterility. However, ulcs1 mutants display several other pleiotropic phenotypes, which are most likely related to the characteristics of the protein to act as a scaffold subunit. As previously suggested, WDR proteins are crucial and indispensable in the assembly of multiple functional complexes that regulate various aspects of plant growth and development. In this report, we present several growth phenotypes associated with the impairment of the AtULCS1 gene and discuss their nature and putative origin in the context of plant development. Furthermore, we show that ULCS1 physically interacts with the UBA-like protein from Arabidopsis thaliana.
Results and Discussion
AtULCS1 encodes for a conserved Cul4-RING E3 ubiquitin ligase subunit
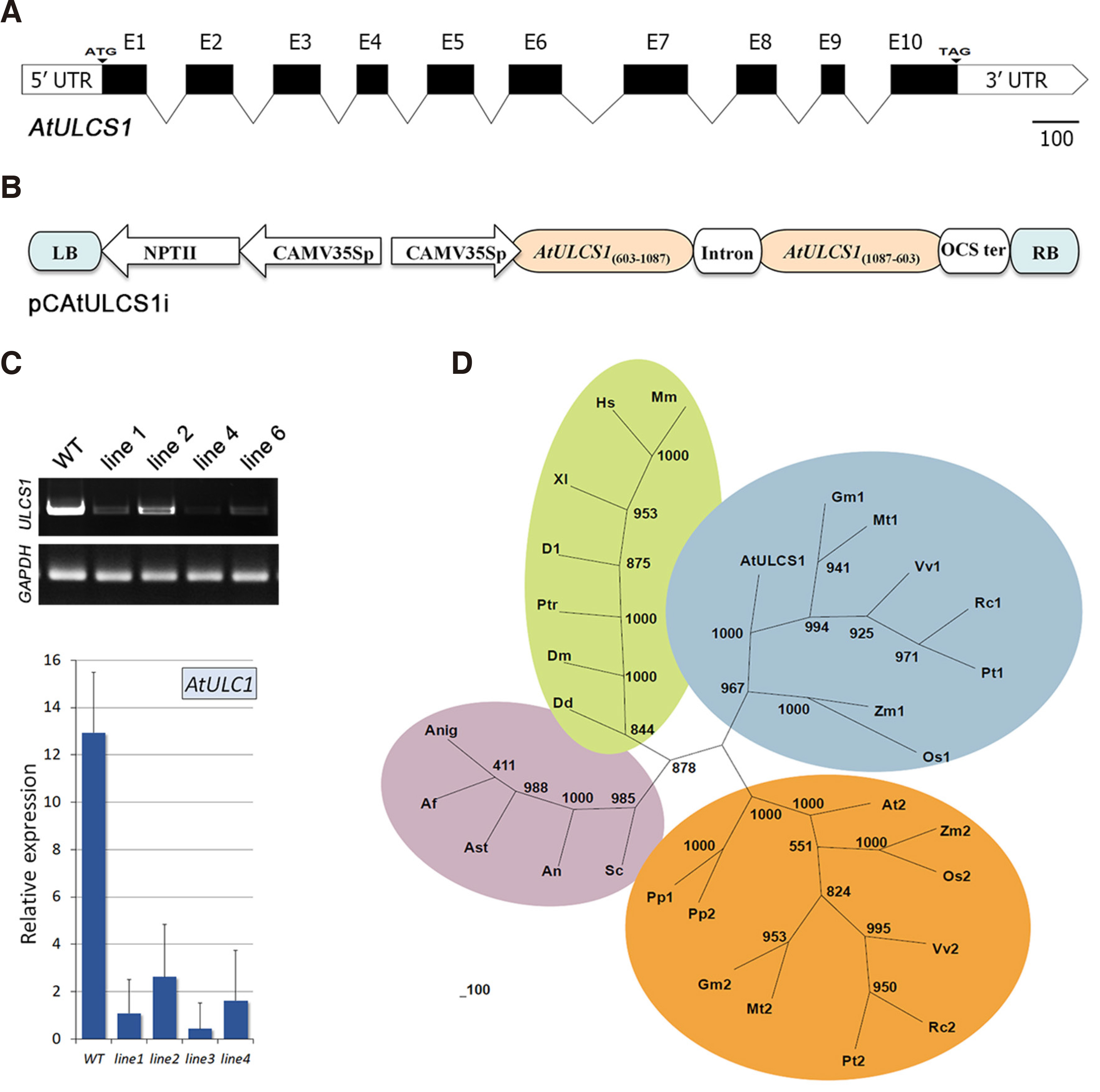
AtULCS1 was previously characterized as a highly conserved WD40-repeat-containing protein of 331 amino acids (Beris et al., 2016). The gene is located on the fifth chromosome of the Arabidopsis thaliana genome at gene locus 66240 and consists of 10 exons (Fig. 1A). The single WD40 repeat-containing domain of AtULCS1 contains four WD40 repeats that cover almost the entire protein sequence, which exhibits a 41-84% amino acid identity across the plant, animal, and fungi kingdom. Almost all plant species contain two closely related homologs with high amino acid identity, which, however, appear to have distinct functions. In Arabidopsis, the two homologs (ULCS1 and APRF1) are shown to be involved in the assembly of a Cul4-RING E3 ubiquitin ligase and a SET1 COMPASS histone methylation complex, respectively (Beris et. al., 2016; Kapolas et al., 2016). The distinct function of the two homologous proteins is underscored by a number of differences found in their amino acid sequence (Beris et. al., 2016). Apart from the characteristic WDxR binding motif, which is present in all orthologs, AtULCS1 contains a unique cysteine instead of glycine (at position 34), while a methionine instead of leucine (at position 48) is present only in Arabidopsis and the respective Zea mays ortholog. Cysteine and methionine amino acids are known to play critical roles in peptide folding and thus protein function. Methionine is one of the most hydrophobic amino acids and tends to participate in the formation of interior hydrophobic protein cores, while oxidation of this amino acid residue within proteins may lead to alterations in protein structure and function. Similarly, cysteine amino acids play vital roles in the formation of tertiary and quaternary structures of proteins, due to their ability to form inter- and intrachain disulfide bonds, thus determining the correct folding and function of the proteins (Brosnan and Brosnan, 2006; Valley et al., 2012). The diverge function of the two homologous sequences present in most plant proteomes is also imprinted in the phylogenetic tree that shows the evolutionary relationship of the orthologs from various eukaryotic species. The analysis showed that AtULCS1 is clustered with other WD40 plant proteins in a distinct clade, formed by the fungi and animal orthologs. In contrast to animals and fungi, which contain only one homolog, the two homologous sequences present in the genome of plant species are also clustered in separate clades of the tree, indicating a distinct functional specification of the two homologs (Fig. 1D).
Fig. 1. Bioinformatic and phylogenetic analysis of ULCS1 (At5g66240) from Arabidopsis thaliana.
(A) Schematic presentation of the AtULCS1 gene structure. (B) Schematic representation of the hairpin RNAi silencing construct (pCAtULCS1i) used to generate impaired transgenic plants. (C) Semi-quantitative RT-PCR (upper panel) and qPCR (lower panel) analysis showing down-regulation of the AtULCS1 transcripts in four independent transgenic silencing lines. Lines ulcs1 1, ulcs1 4, and ulcs1 6 displayed the lowest mRNA levels compared to WT plants. Values ± SD were normalized to GAPDH and represent the mean of three biological samples analyzed in triplicates. (D) Phylogenetic analysis of AtULCS1 orthologous proteins from various species and taxa. Trees were constructed with the neighbor-joining method of PHYLIP after Clustal Omega alignment of the full-length protein sequences. Numbers indicate bootstrap support values from 1,000 replicates. Abbreviations and the corresponding accession numbers are given in Supplementary Table S1.
To investigate the functional involvement of the ULCS1 protein in plant development, we generated a hairpin construct to obtain A. thaliana RNAi-mediated post-transcriptional silencing mutants that were impaired in AtULCS1 function (Fig. 1B). RT PCR analysis performed in individual transgenic T1 and T2 progeny lines showed that AtULCS1 transcripts were significantly downregulated in several of them. Fig. 1C shows four representative lines, displaying moderate and low levels of AtULCS1 mRNA. Seeds from lines 1 and 4 were selected for further phenotypic analysis.
AtULCS1 impaired plants display pleiotropic phenotypes during plant growth and development
We have previously shown that ulcs1 silencing lines exhibited a predominant phenotype of infertility and that the degree of sterility is correlated with the AtULCS1 transcript levels. The analysis revealed that infertility resulted from the inability of the transgenic ulcs1i plants to accomplish secondary wall modifications. IRREGULAR XYLEM gene expression and lignification were significantly diminished, in both the stem vascular tissues and the anther endothecium cells of the silenced plants, leading to an impairment of anther dehiscence and pollen release (Beris et al., 2016). Depending on the severity of impairment, ulcs1 mutants exhibited also several other phenotypic abnormalities, such as increased growth, deformed leaves, and variable seed size and morphology.
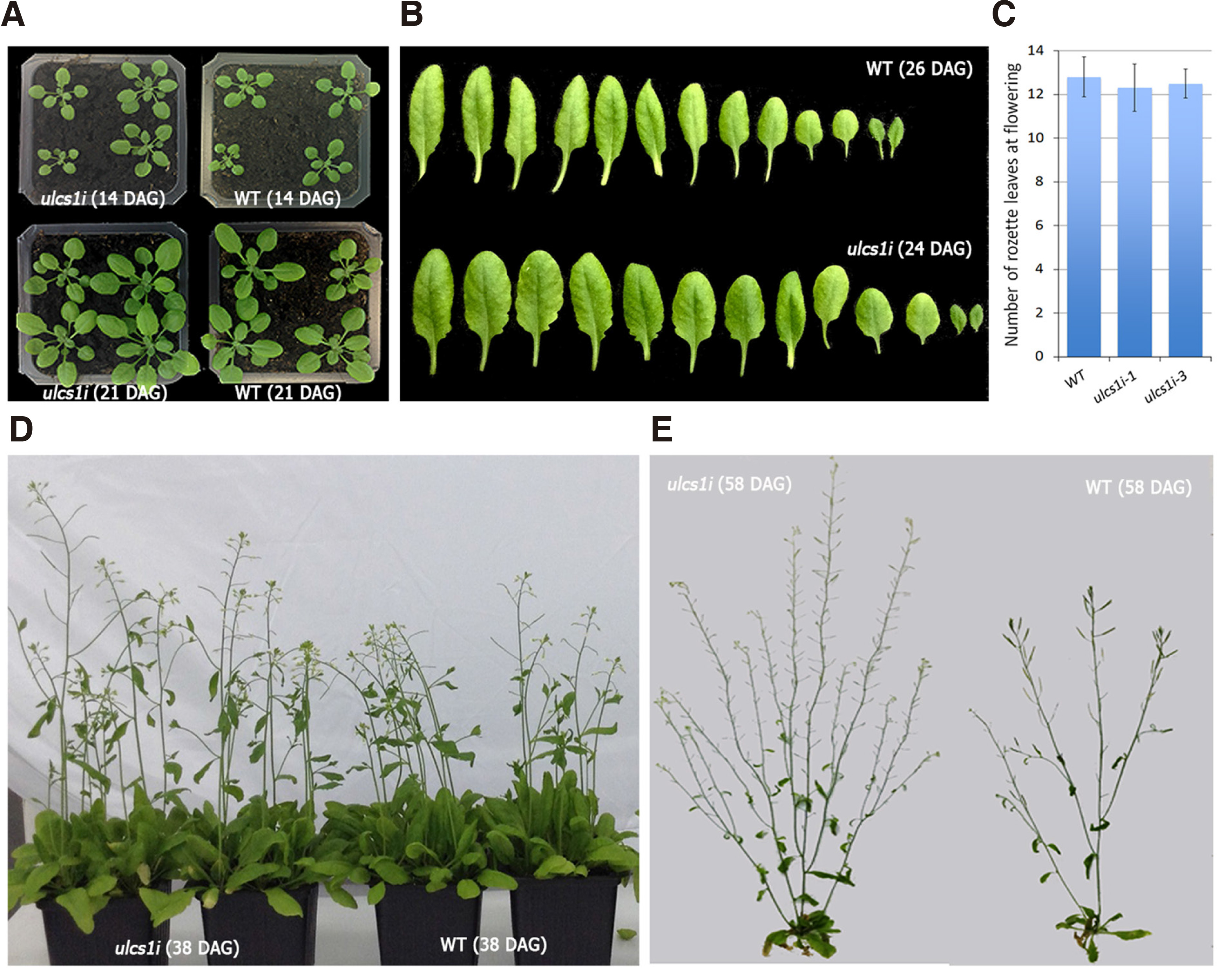
In Arabidopsis, the aerial architecture of the stem, including lateral branching, is established before primary stem bolting (Pouteau and Albertin, 2009). ULCS1 impaired plants exhibited faster development, leading to several vegetative growth alterations. While at 14-days after germination (DAG) WT plants harbored ca. 6 leaves, ulcs1i mutants contained 7-8 leaves. At 21 DAG, the number of leaves was ca. 8 in WT and 10-11 in the ulcs1i plants (Fig. 2A). Rosette leaves of ulcs1i plants were shorter but displayed slightly extended petioles and an increased surface area (Fig. 2B). In Arabidopsis, the transition from the vegetative to the reproductive phase is marked by the emergence of the first flower, followed by bolting transition that is characterized by the elongation of the first internode (Pouteau and Albertin, 2009). Due to accelerated growth, ulcs1i plants reach their vegetative to reproductive transition point earlier than WT plants. (Fig. 2D). However, the emergence of the first flower and internode in ulcs1i plants does not necessarily indicate an early flowering time phenotype. Since these developmental transitions are under the influence of various endogenous and exogenous factors, to evaluate whether a mutation results in an early flowering time phenotype, plants must also be scored based on two other specific indicators, namely the rosette leaf number (RLN) and the number of primary inflorescence internodes at the time of bolting (Pouteau and Albertin, 2011). In WT, the reproductive phase transition initiates at ca. 26 DAG while the plants possess 12-13 rosette leaves. Although ulcs1i plants initiated their reproductive phase at ca. 23-24 DAG, their number of rosette leaves was similar to that of WT plants (Fig. 2C). Furthermore, and as the number of primary inflorescence internodes is concerned, WT plants usually acquire, under long-day light conditions, two to three inflorescence internodes. As shown in Fig. 2D, ulcs1i plants display the same number of internodes during bolting, suggesting that under long-day photoperiod conditions, soil-grown ulcs1i mutants exhibited a similar flowering time phenotype to WT plants. Taken together, the above data clearly indicate that ulcs1i plants may reach their reproductive transition point 2-3 days earlier than WT plants, but this can obviously be attributed to their accelerated growth and, thus, cannot be characterized as early flowering time mutants. However, due to their increased growth rate, ulcs1i plants are at full maturity significantly higher than WT plants and contain an increased number of secondary and lateral branches (Fig. 2E).
Fig. 2. Representative phenotypic analysis of ulcs1i silencing lines during development.
(A) Phenotypes of 14- and 21-day old seedlings of ulcs1i and WT plants (B and C) Comparative depiction of the size, shape, and number of rosette leaves developed in WT and ulcs1i plants, at the time of bolting, 26 days after germination (DAG) for WT and 23-24 DAG for ulcs1i plants. Representative phenotypes of WT and ulcs1 silencing plants at 38 DAG (D) and at full maturity 58 DAG (E).
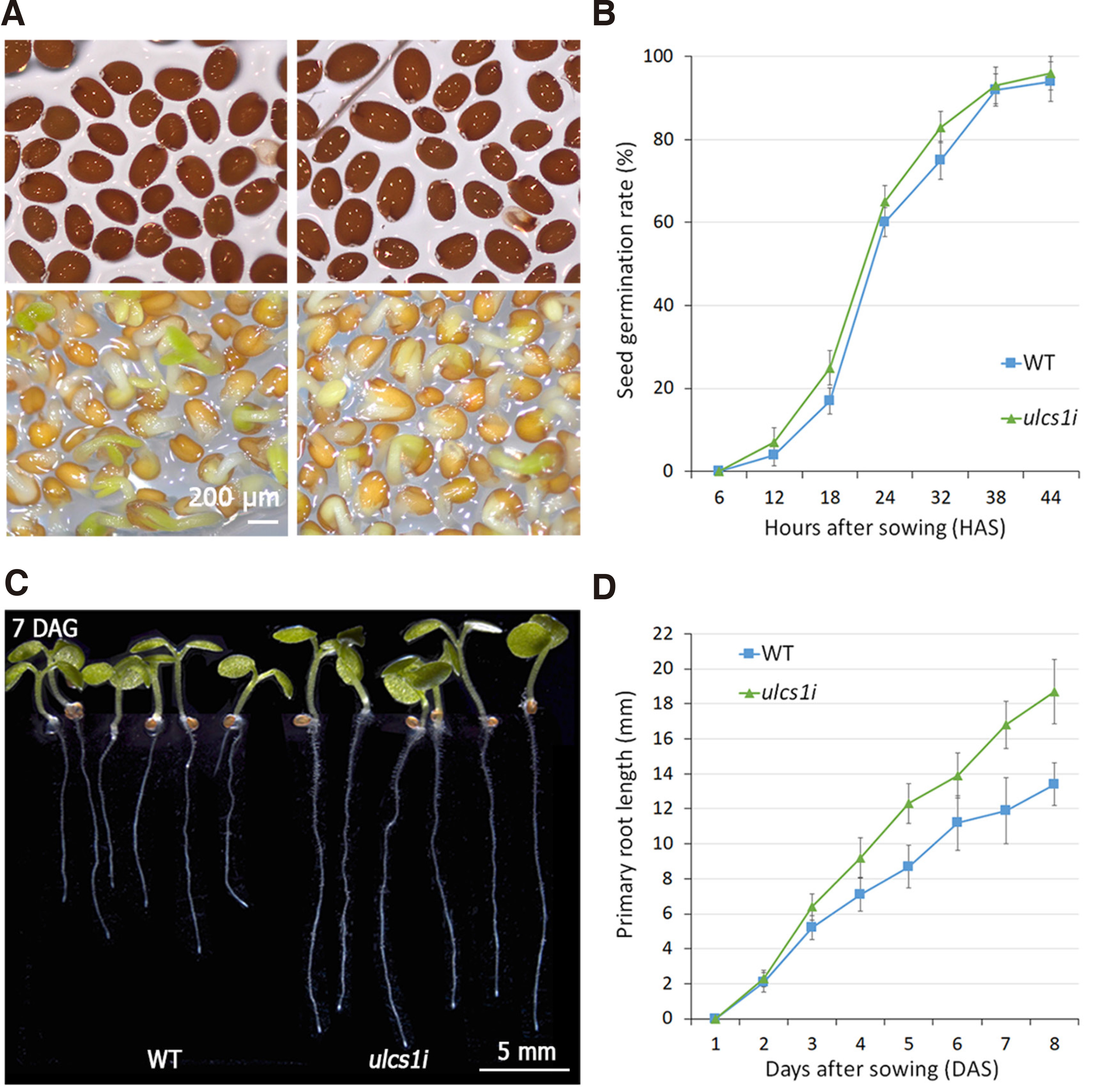
To further investigate the overall faster growth phenotype of the ulcs1i plants, we evaluated seed morphology, the germination rate, as well as the primary root length of WT and mutant plants during seedling development at 1-8 days after sowing. As shown in Fig. 3 A,B, while ulcs1 seeds displayed a greater amplitude of size, compared to the relatively uniform size of WT seeds, they both displayed similar germination curves. Although ulcs1i seeds showed a slightly faster germination rate (by ca. 3-4 hours), compared to WT, their germination curves were almost identical (Fig. 3B). However, after germination, ulcs1i seedlings displayed a constant accelerating growth rate, compared to WT plants, which was statistically significant and evident 7 days after sowing (Fig. 3 C,D).
Fig. 3. Seed germination and seedling growth of WT and ulcs1i silencing lines.
(A) Images showing the size amplitude of imbibed seeds (upper panel) and germinated seeds (lower panel) of WT and an ulcs1i representative line. (B) Seed germination assays showing the germination rate of WT and ulcs1i seeds, HAS: hours after sowing. (C) Representative phenotypes of plants germinated and grown for 7 days on MS medium. (D) Chart showing the primary root length of WT and ulcs1i seedlings grown on MS medium for 1-8 days after sowing (DAS). Data are means ±SE of three biological replicates (n=100 for B and n=20 for D). P value < 0.05 (Student’s t-test).
Mutations in genes encoding for WDR proteins have previously been shown to result in pleiotropic phenotypes. Underexpression of the Solanum chacoense NOTCHLESS (ScNLE) gene, which is essential for cellular growth and proliferation, affected organ number and size, flowering, and seed development (Chantha and Matton, 2007). Similarly, dcaf1 co-suppression mutants exhibit pleiotropic phenotypes such as multiple primary shoots and irregular branching, stunted growth, small rosette leaves, and flower abnormalities (Zhang et al., 2008). On the other hand, bouquet-1 (boq-1), a mis-sense allele of the vernalization independence 3 gene (VIP3) possesses highly pleiotropic shoot growth phenotypes, predominantly caused by an enhancement of meristem formation. The boq-1 plants show an increase in the inflorescence stem number accompanied by frequent fasciation, due to the development of extra SAMs (Takagi and Ueguchi, 2012).
AtULCS1 impaired plants display increased mitotic divisions of meristematic cells
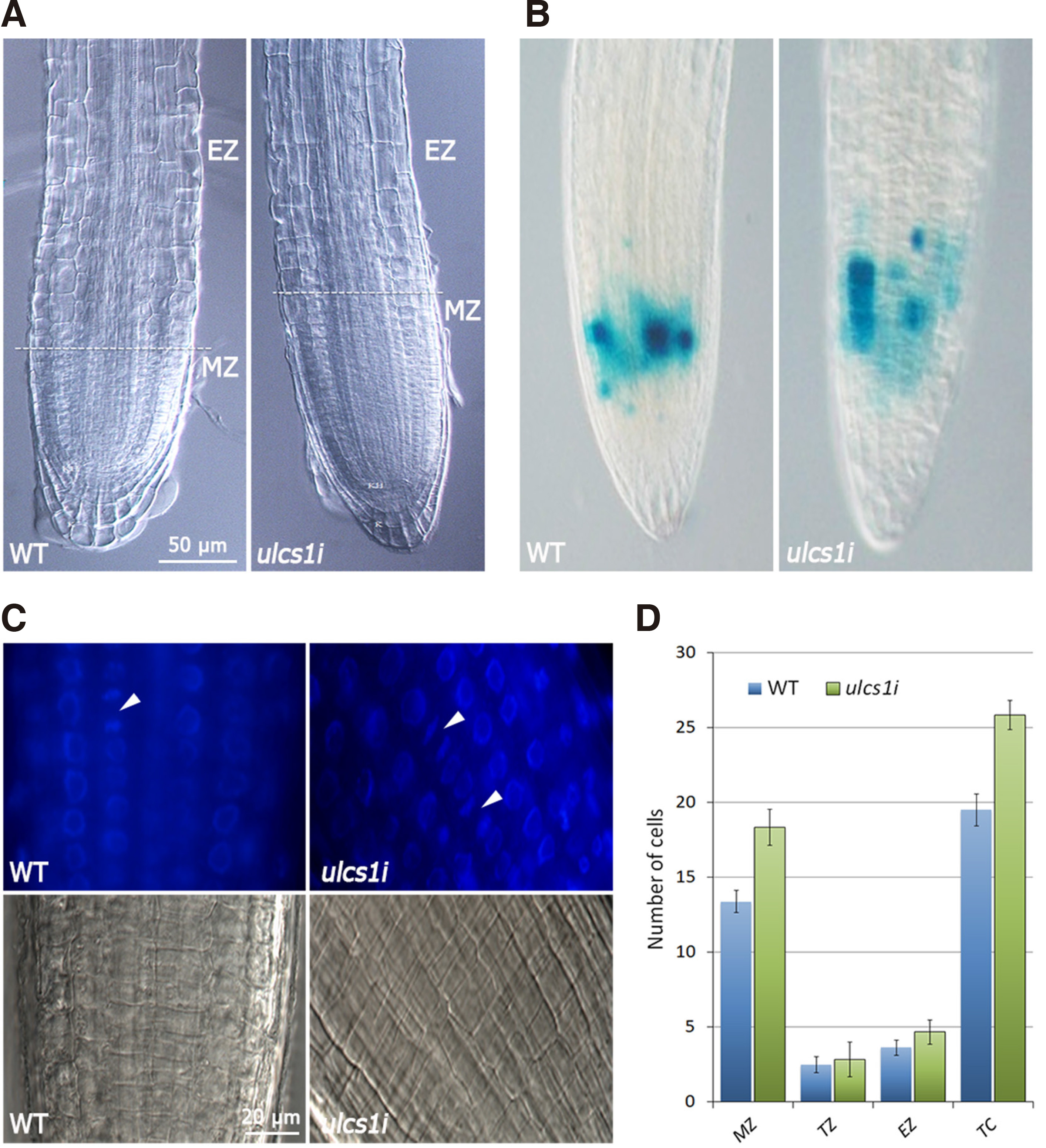
Since the ulcs1i silencing plants were not early flowering mutants, and given the similar seed germination curve compared to the WT plants, we asked whether the accelerated growth was the result of more mitotic divisions that lead to an increased accumulation of cells in meristematic tissues, such as the root apical meristem. To evaluate this hypothesis, we observed roots of ulcs1i and WT plants under a DIC microscope and counted the number of cells of the major root zones at various time intervals during seedling growth. As shown in Fig 4, the meristematic zone of ulcs1i plants was significantly elongated compared with that of WT plants. At five days after sowing, the meristematic zone of WT plants had a length of about 75-80 μm and contained ca. 13 cells, while the corresponding zone of ulcs1i plants was about 120 μm in length and consisted of ca. 18 cells (Fig. 4 A,D). As anticipated, besides the meristematic zone, both the transition and the elongation zone of ulcs1i roots also contained an increased number of cells. Altogether, the root tip of ulcs1i plants, and up to the differentiation zone, consisted of more than 26 cells, compared to the WT root tip, which harbored in total fewer than 20 cells (Fig. 4D). Furthermore, we performed reciprocal crosses between ulcs1i plants and a CYCB1::GUS line, to visualize the number of cell divisions of meristematic cells. GUS staining of the F2 and F3 progeny of plants resulting from the crosses showed that the ulcs1i root meristems exhibited an increased number of mitotic cell divisions, compared with WT (Fig. 4B). This was also evident when the meristems of ulcs1i and WT roots were stained with Hoechst 33258 and observed under DIC and fluorescent microscopy (Fig. 4C).
Fig. 4. Rate of root growth in WT and ulcs1i silencing plants.
(A) DIC images showing root tip of five DAS WT and ulcs1i silencing plants. The horizontal dotted line marks the boundaries between the meristematic and elongation zone. (B) Root-tip of 5-day-old seedlings obtained from crosses between ulcs1i and CYCB1::GUS plants showing GUS-stained meristematic cells undergoing mitotic divisions. (C) Hoechst 33258 stained fluorescent (upper panel) and DIC (lower panel) images of WT and ulcs1i meristematic zones. Arrowheads indicate sites of mitotic divisions of meristematic cells. (D) Chart showing the number of cells present in the different zones of 5-day-old WT and ulcs1i roots. MZ: meristematic zone; TZ: transition zone; EZ: elongation zone; TC: total cells. Data are means ±SE of three biological replicates (n=20). P value < 0.05 (Student’s t-test).
Progression through the cell cycle is an essential step for eukaryotic cells. The SCF type Cullin Ring E3 Ubiquitin Ligases (CRLs) have previously been shown to play a critical role in cell cycle regulation both in animals and plants. They are involved in the degradation of negative regulators, such as cell cycle-dependent kinase inhibitors (CKIs) at the G1 to S phase checkpoint (Zhang and Zhang, 2015). In Arabidopsis, for example, the F-Box Protein FBL17 has been shown to be essential for progression through the second mitosis during pollen development (Kim et al., 2008; Gusti et al., 2009), while a CULLIN 4-RING FINGER-LIGASE complex has been found to play a key role in the control of endoreplication cycles of trichomes (Roodbarkelari et al., 2010). Our results indicate that ULCS1 may participate in a DWD E3 CRL complex that degrades positive regulators of the cell cycle progression machinery.
ulcs1i seeds release larger “halos” upon hydration in water and contain more methylesterified pectins
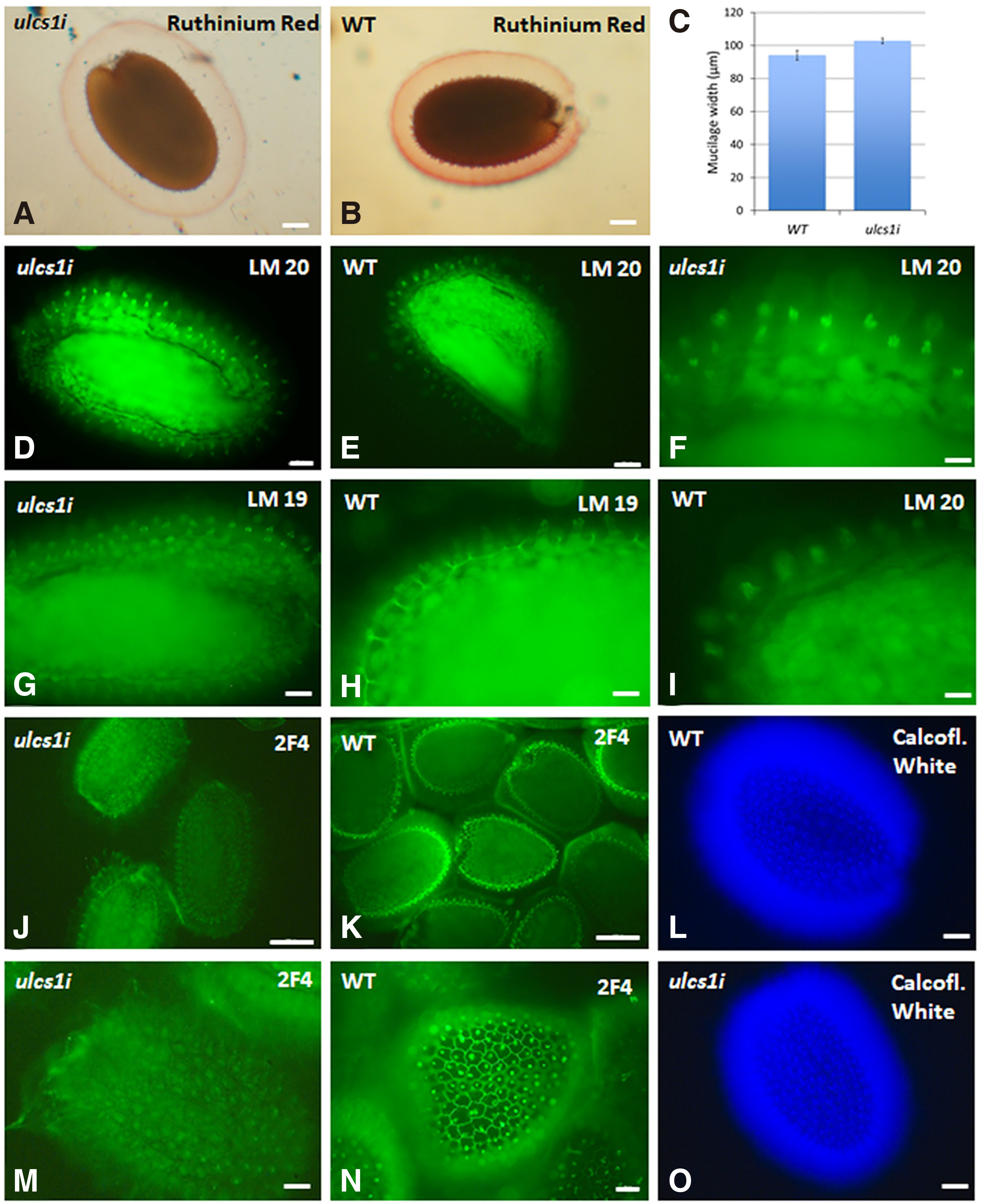
In comparison with WT, hydrated ulcs1i seeds appear to release larger mucilage “halos” (Fig. 5 A,B). The “halos” formed by the adherent layer of the mucilage were around 10 15% wider in ulcs1i seeds than the ones formed around the WT seeds (Fig. 5C). Both “halos” displayed a darker disc at the edge of the mucilage “halo”, as observed by Ruthinium Red staining.
Fig. 5. Ruthenium Red staining and immunolabeling of pectin in seeds and mucilage of WT and ulcs1i silencing plants.
(A and B) Ruthenium Red labeling of ulcs1i (A) and WT (B) “halos”. (C) Measurement of mucilage width of the “halo” after RR staining. Data are means ±SE of three biological replicates (n=60). P value < 0.05 (Student’s t-test). (D, E, F, I) LM20 immunolabeling of heavily methylesterified HGs in ulcs1i (D, F) and WT (E, I) seeds and mucilage. (G, H) LM19 immunolabeling of partially demethylesterified HGs in ulcs1i (G) and WT (H) seeds and mucilage. (J, K, M, N) 2F4 immunolabeling of partially demethylesterified HGs crosslinked by calcium in ulcs1i (J, M) and WT (K, N) seeds and mucilage. (L, O) Calcofluor white staining of WT (L) and ulcs1i (O) seeds. Scale bars in J, K= 200 μm, in A, B, D, E, G, H, M, N=50 μm, and in F, I= 20 μm.
Homogalacturonans (HGAs) are composed of 1,4 linked a-D-galactosyluronic acid residues. Some of their carboxyl groups are methyl esterified. LM20, LM19, and 2F4 monoclonal antibodies recognize different patterns of methyl esterification on HGAs; the LM19 epitope contains low methyl esters, whereas the LM20 epitope is more heavily methyl esterified (Verhertbruggen et al., 2009). The 2F4 monoclonal antibody recognizes non esterified or de-esterified HGAs that are cross-linked by calcium (Sobry et al., 2005; Baluška et al., 2005). Immunolabeling revealed that ulcs1i seeds displayed an intense LM20 fluorescent signal at the mucilage released in rays associated with the tops of the columella of the seed coat epidermal cells (Fig. 5 D,F), in comparison with that observed at the cells of the WT seeds (Fig. 5 E,I). Labeling with LM19 revealed a signal at the radial cell wall of the WT epidermal cells (Fig. 5H), while in ulcs1i, the signal was observed at the mucilage present at the top of the columella of the seed coat epidermal cells (Fig. 5G). When labeling with 2F4 antibody, WT seeds displayed a fluorescent signal at the outer tangential and the radial cell walls. An intense fluorescent signal was also observed at the tops of the columella of the seed coat epidermal cells (Fig. 5 K,N). On the contrary, most of the ulcs1i seeds did not display 2F4 fluorescent signaling on the cell walls of the seeds or at the mucilage present at the top of the columella cells (Fig. 5 J,M). After calcofluor white staining, no significant differences were observed between WT and ulcs1i seeds (Fig. 5 L,O).
The carboxyl groups of HGAs are initially methylesterified in the Golgi apparatus and transported to the plasma membrane for excretion. In the apoplast, HGAs are demethylesterified by the combined action of methylesterases and methylesterase inhibitors (Sola et al., 2019). Methyltransferases needed for the synthesis of mucilage HG have not been identified as yet, but a putative pectin methylesterase, PECTIN METHYLESTERASE 58 (PME58; Turbant et al., 2016), and two putative pectin methylesterase inhibitors, PECTIN METHYLESTERASE INHIBITOR 6 (PMEI6; Saez-Aguayo et al., 2017) and PMEI14 (Shi et al., 2018), are essential in establishing the typical degree of methylesterification (DM) in seed mucilage. Since mutations in PME58 result in modest increases in the DM of seed mucilage, and evidence suggests that PMEI6 does not act on PME58, it is likely that other PMEs are involved in this process. Like PMEI6, the target of PMEI14 is unknown. The subtilisin-like protease SUBTILISIN-LIKE PROTEASE1.7 (SBT1.7, also known as ARA12; Rautengarten et al., 2008) is also required for normal levels of HGAs methylesterification. Mutations in ARA12/SBT1.7, like those in PMEI6, result in a larger adherent mucilage layer, poorer mucilage extrusion, and higher PME activity in the seed. Therefore, it is hypothesized that ARA12/SBT1.7 modulates PME activity, possibly through protein turnover. It has already been reported that FLYING SAUCER 1, a Ring E3 Ubiquitin Ligase regulates the degree of pectin methylesterification in Arabidopsis seed coat mucilage. Voiniciuc et al., 2013, characterized the flying saucer1 (fly1) mutation, which displayed primary wall detachment, reduced mucilage extrusion, and increased mucilage adherence, probably due to the lower degree of methylesterification in the mucilage. FLY1 encodes a transmembrane protein with a RING-H2 domain that has in vitro E3 ubiquitin ligase activity. However, neither the target of FLY1 nor its exact mode of action has been determined. The fact that the DM of HGAs appears to be regulated by protein degradation through two distinct mechanisms (ARA12/SBT1.7 and FLY1) and by at least one PMEI, indicates that mucilage production and extrusion is highly dependent on the tight regulation of the degree of pectin methylesterification (Sola et al., 2019). It would be very interesting to further elucidate the involvement of a putative E3 ULCS1 CRL Ligase complex in the regulation of pectins demethylesterification processes during seed coat development, given that they seem to play a crucial role in mucilage formation and seed germination.
AtULCS1 interacts physically with the UBA-like protein
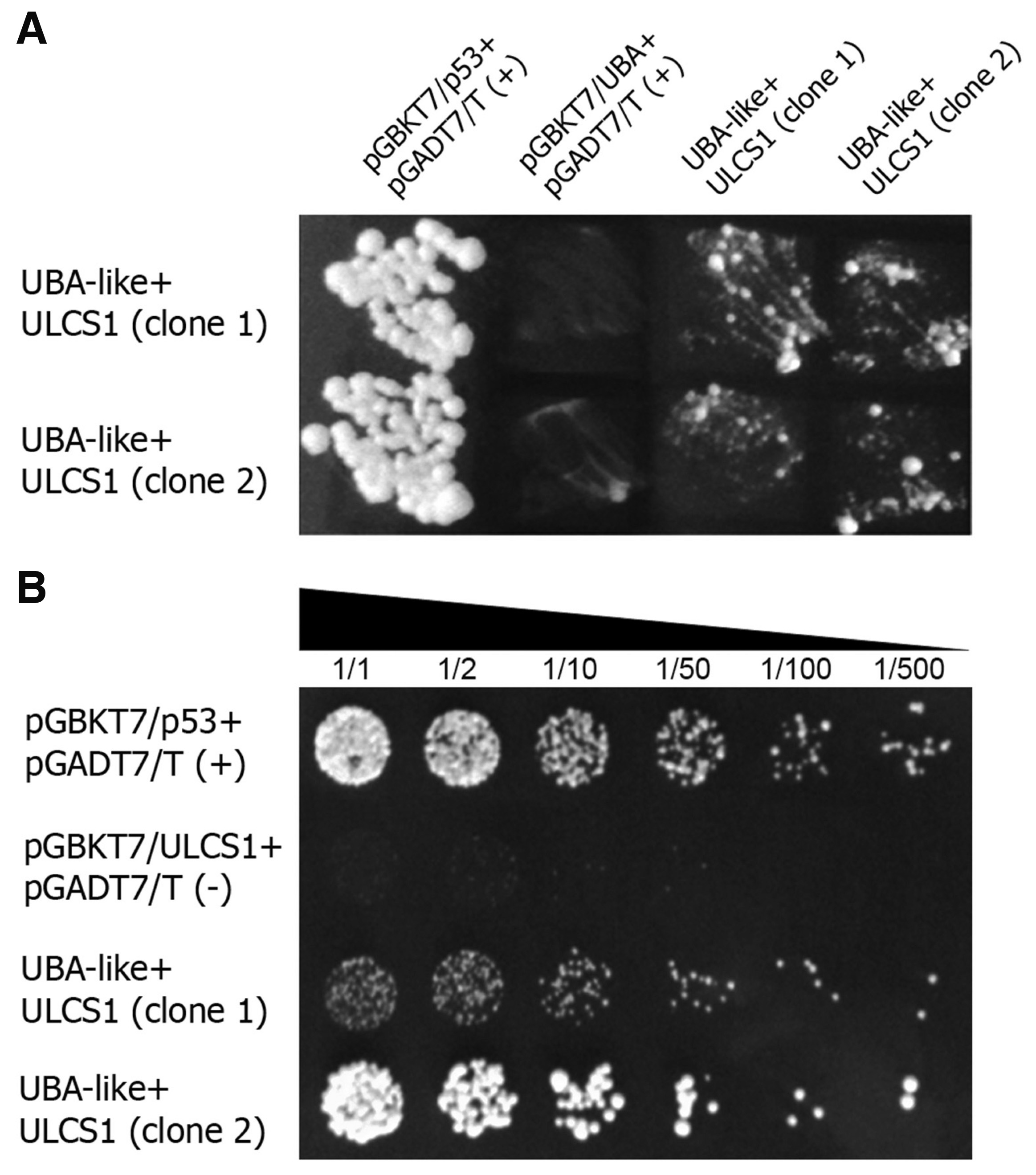
ULCS1 interacts with DDB1a in planta (Beris et al., 2016), suggesting a direct or indirect physical coupling of these proteins and the potential existence of a CRLDDB1a/ULCS1 E3 ligase complex in plants (Beris et al., 2016). Here we show that ULCS1 also interacts physically with the UBA-like protein from Arabidopsis. UBA-like is a basic salivary proline-rich-like protein that, when impaired, results in plants that display similar phenotypes to those observed in ulsc1i mutants (unpublished data). Both ULCS1 and UBA cDNAs were cloned into the appropriate yeast two-hybrid vectors and assayed for their putative association. Fig. 6 shows two representative independent clones, which showed a positive interaction, as indicated by the ability of the yeast cells to grow on selective media (Fig. 6A). To further verify the interaction, series dilution (1/1 to 1/500) of the two independent clones was spotted on selection plates and allowed to grow for 3-4 days. As shown in Fig. 6B colonies from both clones were visible up to dilution 1/500. As negative controls, yeast cells of strain SG335 were transformed with plasmid pGBKT7/UBA or pGBKT7/ULCS1 along with plasmid pGADT7/T and monitored for growth on SD medium selection plates without Trp and Leu. Even at low dilution (1/1), no colonies were visible (Fig. 6B).
Fig. 6. Visualization of protein-protein interaction using yeast two-hybrid analysis.
(A) Heterodimerization of AtULCS1 and two independent clones of AtUBA-like in yeast cells. (B) Series dilution (1/1 to 1/500) of the two positive clones on selection plates. Double transformed SG335 yeast strain colonies were selected on synthetic dropout (SD) medium without Trp and Leu (SD, trp-, leu-) and restreaked on SD/−Trp/−Leu medium containing 1% raffinose and 2% galactose. As a positive control (+), yeast cells SG335 harboring plasmids pGBKT7/p53 and pGADT7/T were used. As negative controls (-) yeast cells of strain SG335 were transformed with plasmid pGBKT7/UBA (A) or pGBKT7/ULCS1 (B) along with plasmid pGADT7/T and monitored for growth on SD selection plates without Trp and Leu. Yeast growth was evaluated after 3-4 d at 30°C.
Materials and Methods
Plant material and growth conditions
Wild-type (WT) and transgenic RNAi Arabidopsis thaliana (L.) Heynh. (ecotype Col-0) plants were grown under standard conditions at 22°C in ca. 60% humidity with a light/dark cycle of 16 h/8 h and illumination of 110 μE m 2 s 1 PAR, supplied by cool-white fluorescent tungsten tubes (Osram, Berlin, Germany). Seeds from WT and individual T2 or T3 transgenic plants were imbibed at 4°C for 24h and were subsequently surface-sterilized for 2 min with 70% (v/v) ethanol and 5 min with 15% (v/v) sodium hypochlorite containing 0.1% (v/v) Tween 20. After several washes with sterile deionized water, seeds were germinated on half-strength Murashige and Skoog (MS) medium without sucrose or on medium containing 50 mg/L kanamycin under the same growth conditions. Transgenic plants were transferred to soil for further development.
Seed germination assays were carried out in petri dishes with two layers of water saturated filter paper (Whatman). Germination was scored in 6-h time intervals for up to 44 h and the germination frequency (%) was calculated. For seedling root growth measurements, seeds were germinated vertically on half-strength MS plates without sucrose for 1-8 days. Before sowing, all seeds were imbibed at 4 °C for 48 h in darkness. Each experiment was repeated three times using biological replicates.
Bioinformatic and phylogenetic analysis
The DNA sequence of the AtULCS1 gene (At5g66240) was obtained from TAIR (http://www.arabidopsis.org). The full-length amino acid sequences used for the phylogenetic analysis were obtained from NCBI (http://www.ncbi.nlm.nih.gov/pmc/) using the BLASTp software (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Alignment of the protein sequences was performed with the CLUSTAL OMEGA server at EBI (http://www.ebi.ac.uk/). The ExPaSy software suite (http://www.expasy.org/) was used for all routine bioinformatics analyses.
The phylogenetic tree was constructed by using the neighbor-joining method as implemented in the NEIGHBOR program of the PHYLIP package, version 3.65 (Department of Genome Science, University of Washington, Seattle, WA, USA). Amino acid distances were calculated according to the Dayhoff PAM matrix or the Protein Parsimony method (PROTDIST and PROTPARS programs of PHYLIP package, respectively). The statistical significance was tested by bootstrap analysis for 1,000 replicates. The final tree was visualized using TreeView 1.6.6. (http://taxonomy.zoology.gla.ac.uk/rod/treeview.html). The numbers indicate the bootstrap replications (of 1,000) in which the given branching was observed. The protein parsimony method produced trees with essentially identical topologies. Accession numbers and gene model names are provided in Supplementary Table S1.
Nucleic acid extraction, cDNA synthesis and gene expression analysis
DNA and total RNA were extracted from plant tissues using the NucleoSpin® Plant II and NucleoSpin® RNA Plant kits, respectively, according to the manufacturer’s instructions (Macherey Nagel, Düren, Germany). First-strand cDNA synthesis was performed using 1 μg of total RNA as template and the PrimeScript Reverse Transcriptase (Takara-Clontech, Kyoto, Japan). PCR products for cloning were amplified with Phusion® High-Fidelity DNA Polymerase (New England Biolabs, Beverly, MA, USA). For semiquantitative (sq-) and quantitative (q) RT-PCR, Taq DNA polymerase (Invitrogen, Carlsbad, CA, USA) and KAPA SYBR® FAST qPCR Kits (Kapa Biosystems, Woburn, MA, USA) were used, respectively, according to the manufacturer’s instructions. As an internal control for RNA calibration, the transcripts of GAPDH or rDNA were monitored. All experiments were performed in triplicate. All PCR products were separated by electrophoresis on 0.9% agarose gels and visualized under UV light after staining with 80 μg L-1 ethidium bromide. The primers used in this study are listed in Supplementary Table S2.
Construction of vectors for plant transformation and yeast two-hybrid assays
To generate the RNAi silencing construct, the DNA fragment spanning the region between the 608th and 1087th nucleotide (including the 3’ UTR) of the AtULCS1 full-length cDNA was amplified by PCR using primer pairs XbaIULCS1-Fi/ClaIULCS1-Ri and EcoRIULCS1-Fi/KpnIULCS1-Ri. The purified bands were then cloned as a KpnI/EcoRI (sense) and XbaI/ClaI (antisense) fragment into the respective cloning sites of the pHannibal vector. The hairpin construct was subsequently introduced into the binary vector pCambia2200 (Cambia Canberra, Australia) as a PstI/SacI fragment (plasmid pCAtULCS1i) and used to transform Arabidopsis thaliana plants.
For the yeast two-hybrid vectors, the first-strand cDNA, synthesized as described above, was used to amplify via PCR the ULCS1 (At5g66240) and UBA-like (Ubiquitin Associated-like - At5g14540) ORFs. The ExpandTM High Fidelity PCR System (ROCHE) was used for all PCR reactions, according to the manufacturer’s protocol. The amplified ORFs were cloned into the pGADT7 and pGBKT7 yeast two-hybrid vectors (Clontech) as EcoRI-BamHI fragments. The vector constructs were used for the transformation of yeast strain SG335 (MATa trp1-901, leu2-3, 112, ura1-52, his3-200, gal4Δ, gal80Δ, GAL2-ADE2, LYS2:: GAL1-HIS3, met2:: GAL7-lacZ), according to the standard lithium acetate protocol (Clontech). Double transformed yeast colonies were selected on a synthetic dropout (SD) medium without Trp and Leu (SD, trp-, leu-) and restreaked on SD/−Trp/−Leu medium containing 1% raffinose and 2% galactose. Yeast growth was evaluated after 3-4 d at 30ºC.
All constructs were checked for integrity and cloning correctness by restriction enzyme analysis and DNA sequencing.
Plant transformation and histochemical GUS assays
The pCAtULCS1i vector was transformed into Agrobacterium tumefaciens strain GV3101 competent cells by using the general freeze-thaw method, as described by An et al, 1988. The transformed bacteria were then used for the stable transformation of Arabidopsis (Col-0) plants via the floral dip method (Bechtold and Pelletier, 1998).
Histochemical assays for GUS activity were performed using 5-bromo-4-chloro-3-indolyl-β-d-glucuronide (X Gluc) as a substrate. Seedlings or dissected tissues were incubated for 2 h at 37 °C in X Gluc reaction buffer (50 mM sodium phosphate buffer, pH 7.2, 0.5 mM potassium ferrocyanide, 0.5 mM potassium ferricyanide, and 2 mM X-Gluc), dehydrated by a series of ethanol washes, and kept in a solution containing 3.7% (w/v) formaldehyde, 50% (w/v) ethanol, and 5% (w/v) acetic acid at 4° C. Samples were treated with the clearing agent chloral hydrate (5 g chloral hydrate dissolved in 1 ml 30% glycerol) before microscopy.
Ruthenium Red Staining and Calcofluor White Staining
Mature dry seeds were hydrated in distilled water for 24 h and stained with 0.01% (w/v) Ruthenium Red (Sigma-Aldrich) for 30 to 60 min. Subsequently, seeds were rinsed several times with distilled water and imaged using a Zeiss Axioplan microscope (Zeiss, Berlin, Germany). For cellulose staining, seeds were stained for 10 min with Calcofluor White solution (Sigma-Aldrich) and observed with an epifluorescent microscope.
Immunolocalization of homogalacturonans
For immunolabeling of LM20, LM19, and 2F4 homogalacturonans (HGAs) epitopes in seeds, the protocol described by Giannoutsou et al., 2016 was applied. LM20, LM19, and 2F4 (Plant Probes, Leeds, UK) were used as primary antibodies and FITC–conjugated anti-rat and anti-mouse IgG (Sigma) as secondary antibodies, respectively. LM20 and LM19 antibodies were diluted 1:40 in PBS containing 2% (w/v) BSA. For the immunolocalization of 2F4 specifically, seeds were not treated enzymatically and 2F4 antibody was directly applied to the seeds, while its secondary antibody was diluted 1:40 in T/Ca/S buffer (Tris-HCl 20 mM pH 8.2, CaCl2 0.5 mM, NaCl 150 mM). During the immunolabeling procedure with the 2F4 antibody, samples were washed with T/Ca/S buffer, according to the manufacturer’s instructions (Plant Probes, Leeds, UK).
Microscopy
Before microscopy and when appropriate, freshly stained tissues were fixed with acetic acid:ethanol (1:8) buffer for 1 h, washed with 100% and 90% ethanol, 30 min each, and incubated overnight at room temperature in chloral hydrate buffer. Root meristem organization and nuclei were visualized using WT and ulcs1i transgenic RNAi plants stained with Hoechst dye. Seedlings were fixed in 4% (wt/vol) PFA in PEM (50 mM Pipes, 5 mM EGTA, 5 mM MgSO4, pH 6.9, with KOH) for 15 min, followed by washing for 5 min in PEM buffer. Fixed seedlings were washed with phosphate-buffered saline (PBS), stained with 10 μg ml-1 Hoechst 33258, and then washed with PBS buffer three times.
All samples were visualized with a Zeiss Axioscope epifluorescence microscope (Zeiss, Oberkochen, Germany), equipped with a Zeiss Axiocam MRc5 digital camera, a differential interference contrast (DIC) system, and proper filters. In particular, a set filter with exciter BP450-490 and barrier BP515-595, a set with exciter G-365 and barrier LP420, and a set with exciter BP510-560 and barrier LP590 were used. Stereoscopic photographs were obtained with a Zeiss Stemi 2000-C stereomicroscope, equipped with Jenoptic ProGres3 digital camera (Jenoptik, Jena, Germany). All images were processed (linear adjustments only) by using Adobe Photoshop CC software (Adobe Systems Inc., San Jose, CA, USA).
Supplementary Material
Acknowledgements
This work was supported by the European Social Fund and National Resources (EPEAK II) PYTHAGORAS and the UoA Special Account for Research Grants (S.A.R.G.). We thank the Salk Institute and the NASC for providing the sequence-indexed Arabidopsis T-DNA insertion lines. The authors also wish to apologize to those authors whose excellent work could not be cited due to space restrictions.
Declarations
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
AJs, Adherens Junctions ; CEDPs, Cardiac and Endothelial dual-progenitor ; CCV, Common cardinal veins ; EGFP, enhanced green fluorescent protein ; ESCs, Embryonic Stem Cells ; FHF, First Heart Field ; KO, Knockout ; LV, Left ventricle ; MAPK, Mitogen-activated protein kinase ; OFT, Outflow tract ; Pvec, VE-cadherin promoter ; SHF, Second Heart Field ; TGF-β, Transforming growth factor-beta ; VEGFR2, Vascular-endothelial growth factor receptor 2 ;References
An G., Edbert P. R., Mitra A., Ha S. B., (1988). Binary vectors. In Plant Molecular Biology Manual. (Ed. Gelvin S. B., Shilperoort R. A., Verma D. P. S., ) Kluwer Academic Publishers, Dordrecht.
Baluška F., Liners F., Hlavačka A., Schlicht M., Van Cutsem P., McCurdy D. W., Menzel D. (2005). Cell wall pectins and xyloglucans are internalized into dividing root cells and accumulate within cell plates during cytokinesis. Protoplasma 225: 141-155.
Bechtold N., Pelletier G., (1998). In Planta Agrobacterium-Mediated Transformation of Adult Arabidopsis thaliana Plants by Vacuum Infiltration. In Arabidopsis Protocols. Methods in Molecular Biology. (Ed. Martinez-Zapater J. M., Salinas J., ) Springer Science and Business Media LLC.
Beris D., Kapolas G., Livanos P., Roussis A., Milioni D., Haralampidis K. (2016). RNAi-mediated silencing of the Arabidopsis thaliana ULCS1 gene, encoding a WDR protein, results in cell wall modification impairment and plant infertility. Plant Science 245: 71-83.
Bjerkan K. N., Jung-Roméo S., Jürgens G., Genschik P., Grini P. E. (2012). Arabidopsis WD REPEAT DOMAIN55 Interacts with DNA DAMAGED BINDING PROTEIN1 and Is Required for Apical Patterning in the Embryo . The Plant Cell 24: 1013-1033.
Brosnan J. T., Brosnan M. E. (2006). The Sulfur-Containing Amino Acids: An Overview. The Journal of Nutrition 136: 1636S-1640S.
Chantha S.C., Matton D. P. (2007). Underexpression of the plant NOTCHLESS gene, encoding a WD-repeat protein, causes pleitropic phenotype during plant development. Planta 225: 1107-1120.
Chen H., Shen Y., Tang X., Yu L., Wang J., Guo L., Zhang Y., Zhang H., Feng S., Strickland E., Zheng N., Deng X. W. (2006). Arabidopsis CULLIN4 Forms an E3 Ubiquitin Ligase with RBX1 and the CDD Complex in Mediating Light Control of Development . The Plant Cell 18: 1991-2004.
Chen L., Hellmann H. (2013). Plant E3 Ligases: Flexible Enzymes in a Sessile World. Molecular Plant 6: 1388-1404.
Figueroa P., Gusmaroli G., Serino G., Habashi J., Ma L., Shen Y., Feng S., Bostick M., Callis J., Hellmann H., Deng X. W. (2005). Arabidopsis Has Two Redundant Cullin3 Proteins That Are Essential for Embryo Development and That Interact with RBX1 and BTB Proteins to Form Multisubunit E3 Ubiquitin Ligase Complexes in Vivo. The Plant Cell 17: 1180-1195.
Fong H. K., Hurley J. B., Hopkins R. S., Miake-Lye R., Johnson M. S., Doolittle R. F., Simon M. I. (1986). Repetitive segmental structure of the transducin beta subunit: homology with the CDC4 gene and identification of related mRNAs.. Proceedings of the National Academy of Sciences 83: 2162-2166.
Giannoutsou E., Apostolakos P., Galatis B. (2016). Spatio-temporal diversification of the cell wall matrix materials in the developing stomatal complexes of Zea mays. Planta 244: 1125-1143.
Gusti A., Baumberger N., Nowack M., Pusch S., Eisler H., Potuschak T., De Veylder L., Schnittger A., Genschik P. (2009). The Arabidopsis thaliana F-Box Protein FBL17 Is Essential for Progression through the Second Mitosis during Pollen Development. PLoS ONE 4: e4780.
Hu X.J., Li T., Wang Y., Xiong Y., Wu X.H., Zhang D.L., Ye Z.Q., Wu Y.D. (2017). Prokaryotic and Highly-Repetitive WD40 Proteins: A Systematic Study. Scientific Reports 7: 10585.
Hua Z., Vierstra R. D. (2011). The Cullin-RING Ubiquitin-Protein Ligases. Annual Review of Plant Biology 62: 299-334.
Jain B. P., Pandey S. (2018). WD40 Repeat Proteins: Signalling Scaffold with Diverse Functions. The Protein Journal 37: 391-406.
Jiang D., Kong N. C., Gu X., Li Z., He Y. (2011). Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development. PLoS Genetics 7: e1001330.
Kapolas G., Beris D., Katsareli E., Livanos P., Zografidis A., Roussis A., Milioni D., Haralampidis K. (2016). APRF1 promotes flowering under long days in Arabidopsis thaliana. Plant Science 253: 141-153.
Kaya H., Shibahara K., Taoka K., Iwabuchi M., Stillman B., Araki T. (2001). FASCIATA Genes for Chromatin Assembly Factor-1 in Arabidopsis Maintain the Cellular Organization of Apical Meristems. Cell 104: 131-142.
Kim H. J., Oh S. A., Brownfield L., Hong S. H., Ryu H., Hwang I., Twell D., Nam H. G. (2008). Control of plant germline proliferation by SCFFBL17 degradation of cell cycle inhibitors. Nature 455: 1134-1137.
Lambright D. G., Sondek J., Bohm A., Skiba N. P., Hamm H. E., Sigler P. B. (1996). The 2.0 Å crystal structure of a heterotrimeric G protein. Nature 379: 311-319.
Lechner E., Xie D., Grava S., Pigaglio E., Planchais S., Murray J. A.H., Parmentier Y., Mutterer J., Dubreucq B., Shen W.H., Genschik P. (2002). The AtRbx1 Protein Is Part of Plant SCF Complexes, and Its Down-regulation Causes Severe Growth and Developmental Defects. Journal of Biological Chemistry 277: 50069-50080.
Li Q., Zhao P., Li J., Zhang C., Wang L., Ren Z. (2014). Genome-wide analysis of the WD-repeat protein family in cucumber and Arabidopsis. Molecular Genetics and Genomics 289: 103-124.
Neer E. J., Schmidt C. J., Nambudripad R., Smith T. F. (1994). The ancient regulatory-protein family of WD-repeat proteins. Nature 371: 297-300.
Nelson C. J., Millar A. H. (2015). Protein turnover in plant biology. Nature Plants 1: 15017.
Onouchi H., Igeño M. I., Périlleux C., Graves K., Coupland G. (2000). Mutagenesis of Plants Overexpressing CONSTANS Demonstrates Novel Interactions among Arabidopsis Flowering-Time Genes . The Plant Cell 12: 885-900.
Pouteau S., Albertini C. (2011). An assessment of morphogenetic fluctuation during reproductive phase change in Arabidopsis. Annals of Botany 107: 1017-1027.
Pouteau S., Albertini C. (2009). The significance of bolting and floral transitions as indicators of reproductive phase change in Arabidopsis. Journal of Experimental Botany 60: 3367-3377.
Rautengarten C., Usadel B., Neumetzler L., Hartmann J., Büssis D., Altmann T. (2008). A subtilisin-like serine protease essential for mucilage release from Arabidopsis seed coats. The Plant Journal 54: 466-480.
Roodbarkelari F., Bramsiepe J., Weinl C., Marquardt S., Novák B., Jakoby M. J., Lechner E., Genschik P., Schnittger A. (2010). CULLIN 4-RING FINGER-LIGASE plays a key role in the control of endoreplication cycles in Arabidopsis trichomes. Proceedings of the National Academy of Sciences 107: 15275-15280.
Saez-Aguayo S., Rautengarten C., Temple H., Sanhueza D., Ejsmentewicz T., Sandoval-Ibañez O., Doñas D., Parra-Rojas J. P., Ebert B., Lehner A., Mollet J.C., Dupree P., Scheller H. V., Heazlewood J. L., Reyes F. C., Orellana A. (2017). UUAT1 Is a Golgi-Localized UDP-Uronic Acid Transporter That Modulates the Polysaccharide Composition of Arabidopsis Seed Mucilage. The Plant Cell 29: 129-143.
Scheuring D., Künzl F., Viotti C., Yan M. S.W., Jiang L., Schellmann S., Robinson D. G., Pimpl P. (2012). Ubiquitin initiates sorting of Golgi and plasma membrane proteins into the vacuolar degradation pathway. BMC Plant Biology 12: 164.
Shi D., Ren A., Tang X., Qi G., Xu Z., Chai G., Hu R., Zhou G., Kong Y. (2018). MYB52 Negatively Regulates Pectin Demethylesterification in Seed Coat Mucilage . Plant Physiology 176: 2737-2749.
Smith T. F., Gaitatzes C., Saxena K., Neer E. J. (1999). The WD repeat: a common architecture for diverse functions. Trends in Biochemical Sciences 24: 181-185.
Sobry S., Havelange A., Van Cutsem P. (2005). Immunocytochemistry of pectins in shoot apical meristems: consequences for intercellular adhesion. Protoplasma 225: 15-22.
Šola K., Dean G. H., Haughn G. W., (2019). Arabidopsis Seed Mucilage: A Specialised Extracellular Matrix that Demonstrates the Structure–Function Versatility of Cell Wall Polysaccharides. Wiley 1085-1116.
Stirnimann C. U., Petsalaki E., Russell R. B., Müller C. W. (2010). WD40 proteins propel cellular networks. Trends in Biochemical Sciences 35: 565-574.
Takagi N., Ueguchi C. (2012). Enhancement of meristem formation by bouquet-1 , a mis-sense allele of the VERNALIZATION INDEPENDENCE 3 gene encoding a WD40 repeat protein in Arabidopsis thaliana . Genes to Cells 17: 982-993.
Turbant A., Fournet F., Lequart M., Zabijak L., Pageau K., Bouton S., Van Wuytswinkel O. (2016). PME58 plays a role in pectin distribution during seed coat mucilage extrusion through homogalacturonan modification. Journal of Experimental Botany 67: 2177-2190.
Valley C. C., Cembran A., Perlmutter J. D., Lewis A. K., Labello N. P., Gao J., Sachs J. N. (2012). The Methionine-aromatic Motif Plays a Unique Role in Stabilizing Protein Structure. Journal of Biological Chemistry 287: 34979-34991.
van Nocker S., Ludwig P. (2003). The WD-repeat protein superfamily in Arabidopsis: conservation and divergence in structure and function. BMC Genomics 4: 50.
Verhertbruggen Y., Marcus S. E., Haeger A., Ordaz-Ortiz J. J., Knox J. P. (2009). An extended set of monoclonal antibodies to pectic homogalacturonan. Carbohydrate Research 344: 1858-1862.
Voiniciuc C., Dean G. H., Griffiths J. S., Kirchsteiger K., Hwang Y. T., Gillett A., Dow G., Western T. L., Estelle M., Haughn G. W. (2013). FLYING SAUCER1 Is a Transmembrane RING E3 Ubiquitin Ligase That Regulates the Degree of Pectin Methylesterification in Arabidopsis Seed Mucilage . The Plant Cell 25: 944-959.
Wang W., Zhang Q., Guo D. (2016). An <i>Artemisia</i> WD40-Repeat Gene Regulates Multiple Cellular Functions in Arabidopsis. Journal of Biosciences and Medicines 04: 30-36.
Xu C., Min J. (2011). Structure and function of WD40 domain proteins. Protein & Cell 2: 202-214.
Xu N., Gao X.Q., Zhao X. Y., Zhu D. Z., Zhou L. Z., Zhang X. S. (2011). Arabidopsis AtVPS15 is essential for pollen development and germination through modulating phosphatidylinositol 3-phosphate formation. Plant Molecular Biology 77: 251-260.
Yu L., Smith T. F., Gaitatzes C. (2000). Thirty-plus functional families from a single motif. Protein Science 9: 2470-2476.
Zhang Y., Feng S., Chen F., Chen H., Wang J., McCall C., Xiong Y., Deng X. W. (2008). Arabidopsis DDB1-CUL4 ASSOCIATED FACTOR1 Forms a Nuclear E3 Ubiquitin Ligase with DDB1 and CUL4 That Is Involved in Multiple Plant Developmental Processes . The Plant Cell 20: 1437-1455.
Zhang C., Zhang F. (2015). The Multifunctions of WD40 Proteins in Genome Integrity and Cell Cycle Progression. Journal of Genomics 3: 40-50.